Figures & data
Figure 2. Construction of three Nb expression vectors: pCold-Nb (A), pSUMO-Nb (B) and pMAL-c2X-Nb (C) and PCR amplification (D) of the VHH fragments: M, DNA marker; Lane 1, Nb1 containing a BamHI restriction site; Lane 2, Nb2 containing a SalI restriction site; Lane 3, Nb1 containing a BsaI restriction site; Lane 4, Nb2 containing a BamHI restriction site; Lane C, negative control.
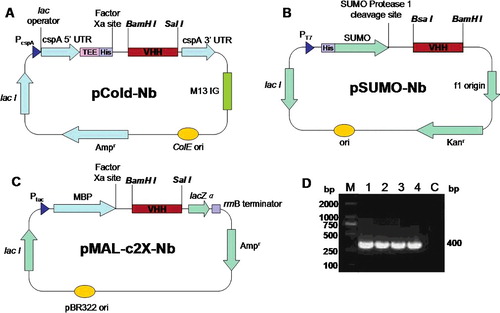
Figure 3. Expression and solubility evaluation of VHH (Nb1 and Nb2) fusion proteins in E. coli: MBP-Nb (55 kDa) in E. coli TB1 (A); His-SUMO-Nb (30 kDa) in E. coli Rosetta (B); His-Nb (20 kDa) in E. coli BL21 (DE3) (C). Arrows indicate the fusion proteins. M, protein marker; IS, supernatant of induced lysates; IP, precipitate of induced lysates; US, supernatant of uninduced lysates; UP, precipitate of uninduced lysates.
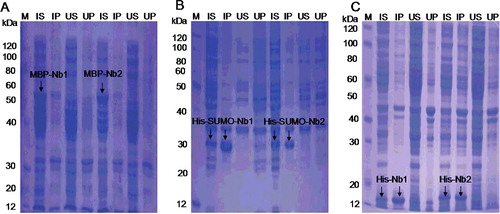
Figure 4. Evaluation of antigen-binding activities of purified MBP-Nbs, His-SUMO-Nbs and His-Nbs using indirect ELISA: Nb1 fusions (A) and Nb2 fusions (B); regression analysis (C) of the antigen-binding activities of purified MBP-Nbs, His-SUMO-Nbs and His-Nbs.
Figure 5. Removal of SUMO tag (A) and purification of native Nb2 (B). (A) Digestion efficiency of SUMO protease 1 (Ulp1) analyzed by 12% SDS-PAGE. (B) Purification of native Nb2. Lane M (A,B), protein maker; Lane 1 (A,B), purified His-SUMO-Nb2 fusion; Lane 2: His-SUMO-Nb2 digested by Ulp1 (A) and purified native Nb2 (B).
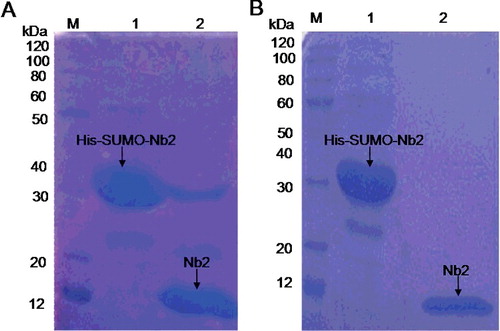
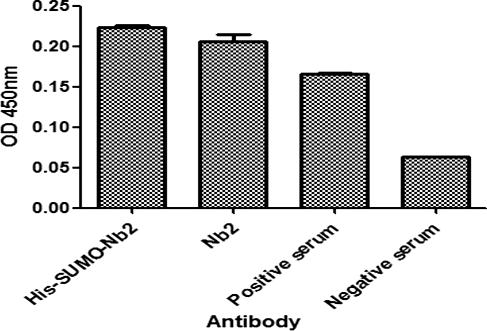
Figure 6. Comparison of the antigen-binding activity of His-SUMO-Nb2 and native Nb2 using double antibody sandwich ELISA. The plates were coated with His-SUMO-Nb2 (2 nmol/L), natural Nb2 (2 nmol/L), positive serum (positive control) and negative serum (negative control). The antigen was a purified native Cap protein, and MBP-Nb1 was added as the second antibody, followed by the addition of HRP-conjugated anti-MBP tag mouse monoclonal antibody.