Figures & data
Figure 1. Construction of pBI–LFchimera-HIS expression vector for LFchimera expression in tobacco. (A) oligonucleotide sequence of native and plant optimized LFchimera. The 6 × His tag and the ER-retention signal KDEL were fused at the C-terminus in-frame with the LFchimera CDS. (B) Plasmid pBI121 contained the nptII (kanamycin-resistance) gene under the control of the nopaline synthase (NOS) gene promoter and a codon-optimized LFchimera under the control of the CaMV 35S promoter and with a NOS terminator. RB, T-DNA right border; LB, T-DNA left border.

Figure 2. Screening transgenic plants. (A) PCR amplification of a 197-bp fragment of LFchimera using DNA derived from N. tobacco plants; Lane M, Ruler 1-kb DNA Ladder Mix (Fermentas, Germany); Lanes 1–5, transgenic plant lines; lane P, positive control (A. tumefaciens culture); Lane C, PCR negative control (water); and Lane NT, non-transgenic plant line. (B) RT-PCR analysis of LFchimera in NT and transgenic plants. The NtEF1a gene was used as a control.
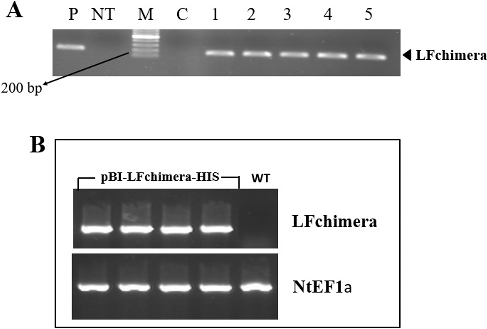
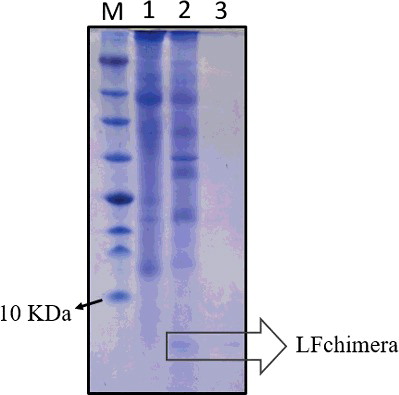
Figure 3. Expression and partial purification of recombinant LFchimera from plant total soluble protein. Tricine SDS-PAGE (16.5%) showing recombinant peptide and purification profile of LFchimera. Lane M, molecular weight marker (Fermentas); Lane 1, total protein from the NT plant; Lane 2, total protein from the T3 transgenic plant; and Lane 3, purified LFchimera using the affinity column.