Figures & data
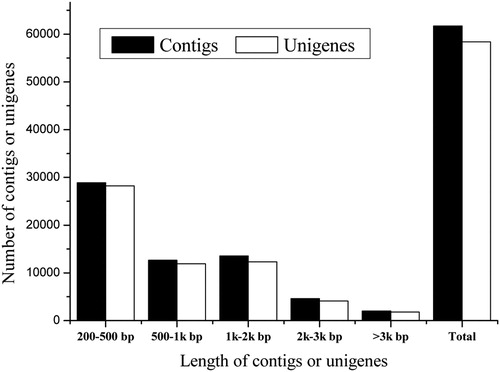
Table 1. Summary of transcriptome sequencing dataset.
Figure 2. Number of unigenes annonated in protein databases by BLASTx with E-value threshold of 10−5. The numbers in the circles indicate the number of unigenes annotated by single or multiple databases.
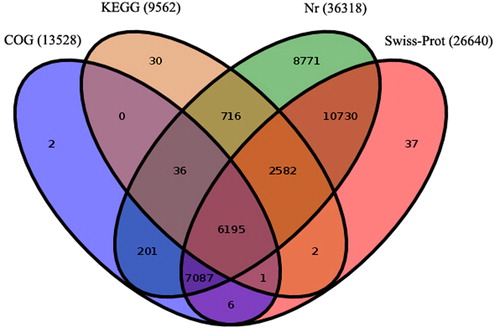
Figure 3. Gene ontology classification of assembled unigenes of Euphorbia kansui. In total 15,506 unigenes were categorized into three main categories: biological process, cellular component and molecular function.
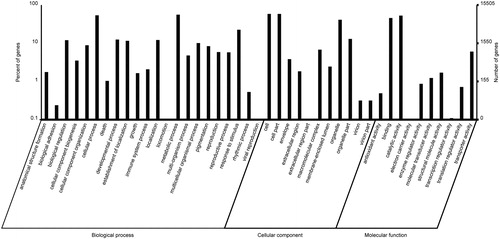
Figure 4. Cluster of Orthologous Groups (COG) function classification of Euphorbia kansui unigenes. A total of 13,528 unigenes were annotated and classified into 25 different classifications.
Figure 5. Terpenoid backbone and diterpene biosynthesis pathways. Terpenoid backbone (isopentenyl diphosphate) is synthesized from two distinct pathways, methylerythritol phosphate (MEP/DXP) pathway and mevalonate (MVA) pathway. The enzymes participating in each step of these pathways are listed in .
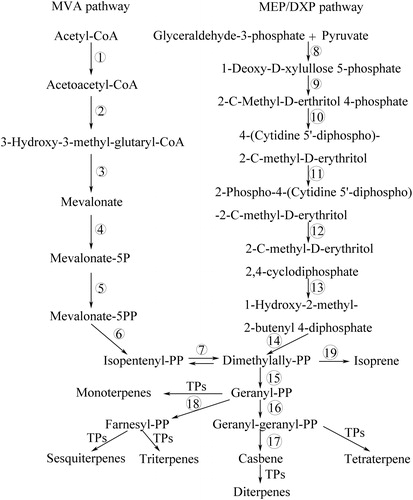
Table 2. Identified unigenes of terpenoid backbone and diterpene biosynthesis.
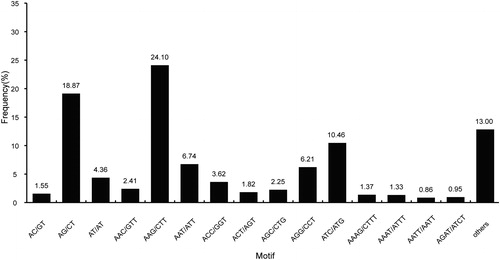
Figure 6. Distribution of simple sequence repeats (SSR) in Euphorbia kansui: distribution of different repeat types (a); number of SSRs per different tandem repeat motifs (b).
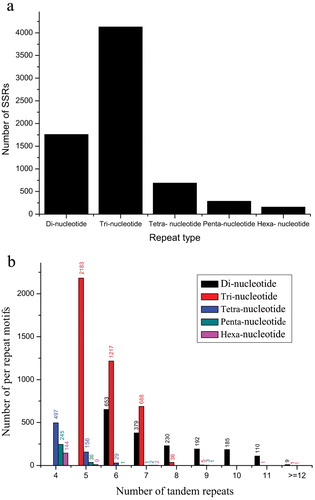
Table 3. Summary of SSR search results.
Table 4. Characterization of 23 polymonophic SSR markers in Euphorbia kansui.