Figures & data
Figure 1. General structures of (a) bisquinoline (b) N1-(7-chloro-4-quinolyl)-1,4-bis(3-aminopropyl)piperazine amide derivatives and (c) N1-(7-chloro-4-quinolyl)-1,4-bis(3-aminopropyl)piperazine amine derivatives as antimalarial agents against the chloroquine-resistant P. falciparum FcB1.
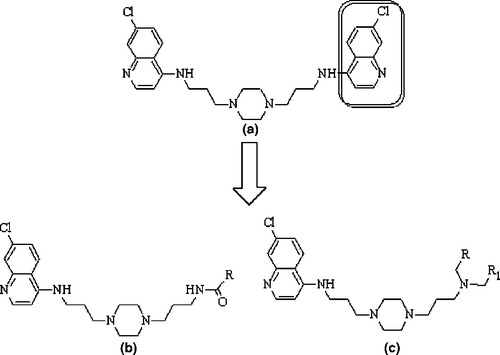
Table I. Observed and modeled in vitro antimalarial activity of N1-(7-chloro-4-quinolyl)1,4-bis(3-aminoprpyl)piperazine- amide derivatives () against the FcB1R strain of P. falciparum.
Table II. Observed and modeled in vitro antimalarial activity of N1-(7-chloro-4-quinolyl)1,4-bis(3-aminoprpyl)piperazine-amine derivatives () against the FcB1R strain of P. falciparum.
Figure 2. Plots of training (▴) and test (ο) sets predicted activities versus observed activity corresponding to Equations (1)(a, b) and (2) (c, d)Equations (1)(a, b) and (2) (c, d). Number of compounds in training set is 21 and in test set is 12.
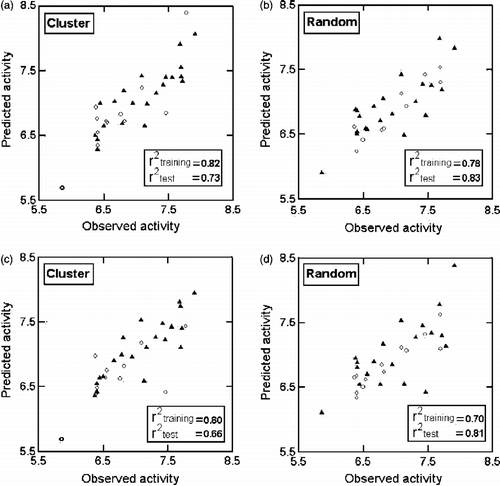
Figure 3. Plot of fraction contribution of MLR-like PLS coefficients (normalized) of the 6 descriptors from Equations (1) and (2) to the activity. The serial numbers 1 to 6 on the horizontal axis refer to the descriptors nDB, ICR, T(n..Cl), GATS1e, GATS2p, and nHDon, respectively.
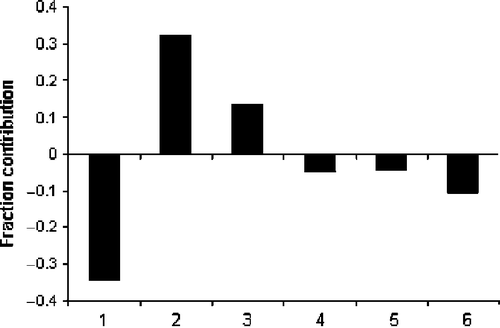
Figure 4. Plots of training (▴) and test (ο) sets predicted activities versus observed activity corresponding to Equations (4) (a, b) and (5) (c, d). Number of compounds in training set is 30 and in test set is 14.
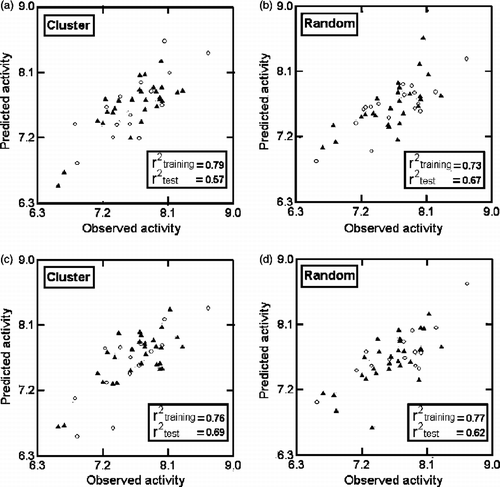
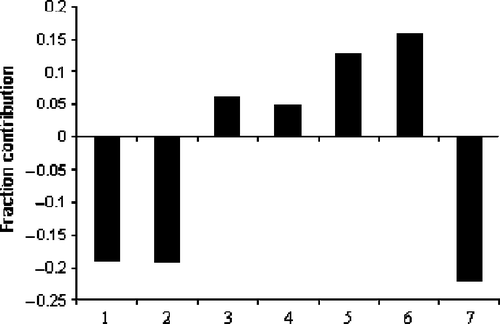
Figure 5. Plot of fraction contribution of MLR-like PLS coefficients (normalized) of the 7 descriptors from Equations (4) and (5) to the activity. The serial numbers 1 to 7 on the horizontal axis refer to the descriptors GMTIV, S0K, ICR, IVDE, MATS8m, MATS6e and Hy, respectively.