Figures & data
Figure 2. Multiple alignment of PDE3A (blue) and PDE3B (green). The residues in the active site of PDE3B (within 15å) are highlighted in yellow (generated by Clustal X (1.81)).

Table 3. Docking analysis data of consensus conformers.
Table 1. Effects of the test compounds upon force of contractility of whole atria from reserpine-pretreated rats: comparison with IBMX, amrinone and cilostamide.
Table 2. Effects of the test compounds upon frequency rate of whole atria from reserpine-treated rats: comparison with IBMX amrinone and cilostamide.
Figure 4. Superimposition of the consensus bonding conformations of cilostamide (black), 4c-d and 4h-m in green stick in the PDE3B active site. (Protein Data Bank: 11 SO)
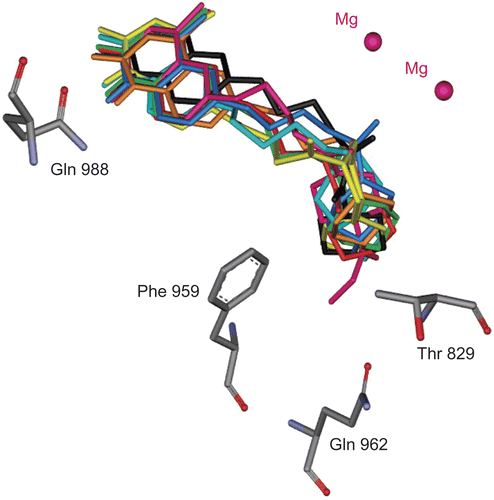
Figure 5. Stick (above) and two view of solvent surface (below) of the interacted amino acids with 4j (colored stick with transparent surface). The three proposed regions A, B and C are distinguished by yellow, green and blue respectively.
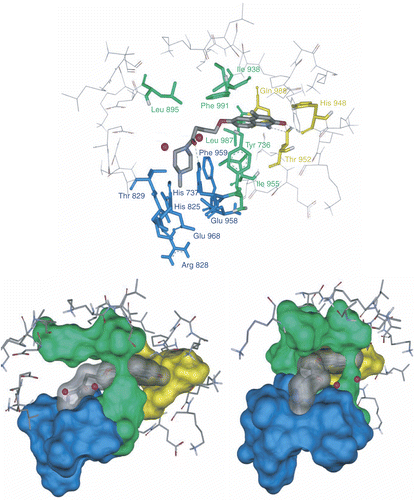
Figure 6. Clustal X (1.81) multiple alignment of human PDEs. The amino acids in the regions A, B and C are highlighted by yellow, green and blue respectively.

Figure 7. Solvent surface (green) of pocket C conserved amino acids having polar and non-polar interactions with consensus structure of compounds 4l (red stick), 4m (green stick), 4j (yellow stick) and 4k (bleu stick). The amide groups of inhibitors have been drawn in contrasted colors.
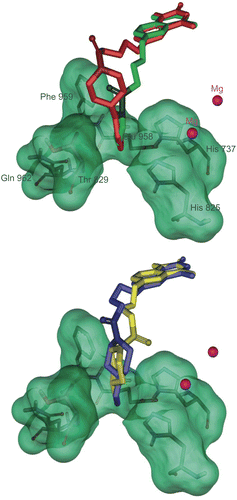
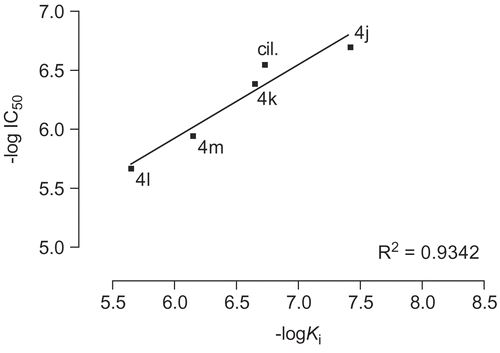
Figure 8. Diagram of increase in force of contractility versus estimated inhibition constant (Ki) for compounds 4c-l.
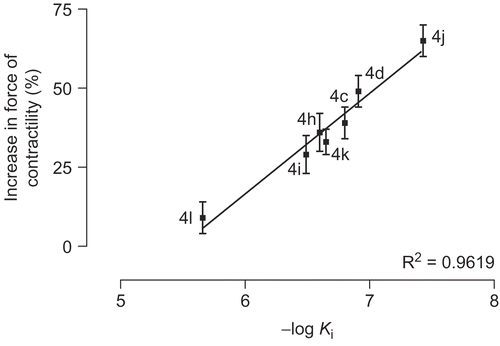
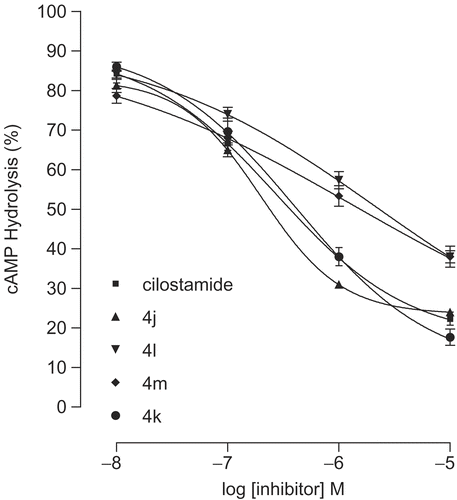
Figure 9. Concentration-effect curves for inhibition of the activity of PDE3 partially purified from rat ventricle by cilostamide and 4j-m. Results are means ± SEM from four experiments. Data are expressed as percent inhibition of the basal enzyme activity (0.35 ± 0.004 pmol hydrolyzed cAMP/μL of protein/min).