Figures & data
Scheme 1. The synthesis of the hydroxamate inhibitors: (a) H-Phe-NHBzl·CF3COOH (for 2a) or H-Leu-NHBzl·CF3COOH (for 3a), TEA, THF; (b) H2, 10% Pd/C, MeOH.
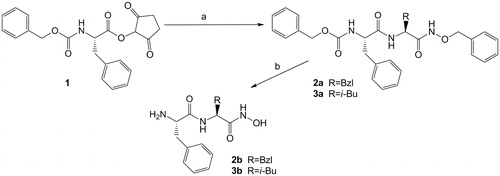
Table 1. Inhibition of human DPP III by dipeptidyl hydroxamic acids at pH 8.0Table Footnote*.
Table 2. Inhibition of human and bacterial DPP III by dipeptidyl hydroxamic acids at pH 7.4Table Footnote*.
Figure 1. Multiple sequence alignment (MSA) of human (UniProt entry: Q9NY33) and B. thetaiotaomicron (UniProt: Q8A6N1) protein sequences. MSA was obtained using ClustalO (www.uniprot.org./align). A section of MSA comprising the constituents of the S1 and S2 subsite, marked with 1 and 2, below alignment is presented in the figure. Five evolutionary conserved regions of the M49 family are framed. Identical amino acid constitutents of S1 and S2 subsites are shadowed grey and those which differ are printed in white on black.

