Figures & data
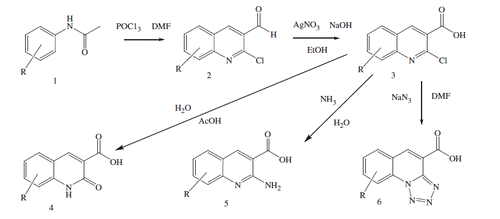
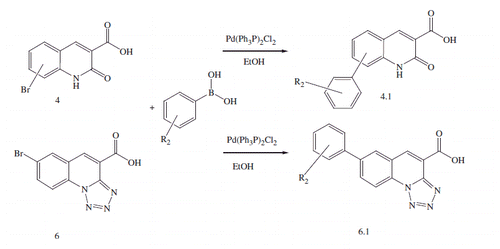
Scheme 3. Synthesis of 5-oxo-2,3-dihydro-1H,5H-pyrido[3,2,1-ij]quinoline-6-carboxylic acid derivatives.
![Scheme 3. Synthesis of 5-oxo-2,3-dihydro-1H,5H-pyrido[3,2,1-ij]quinoline-6-carboxylic acid derivatives.](/cms/asset/6f573c61-b721-42e2-95cd-4c490daea4e9/ienz_a_1222584_sch0003.gif)
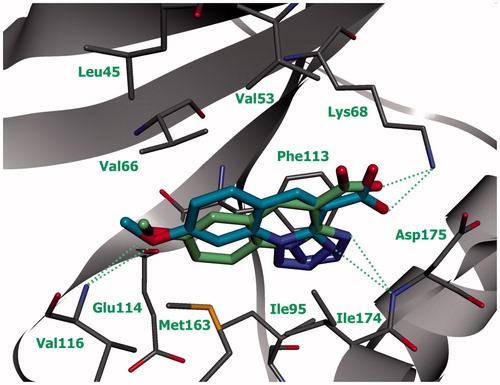
Figure 1. Schematic representation of the complex of compound 93 with active site of protein kinase CK2 (green line indicates amino acid residues involved in hydrophobic contacts; dotted lines indicate hydrogen bonds).
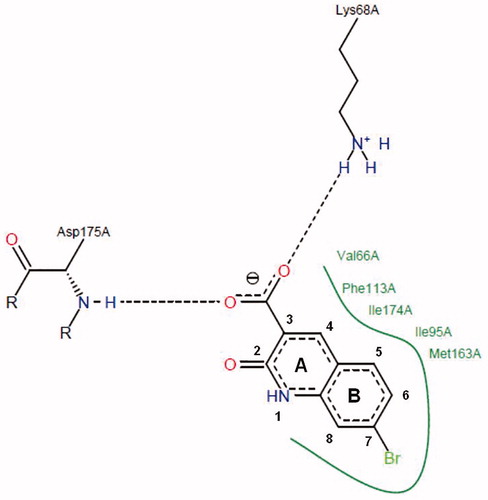
Table 1. Structure of 2-chloro- and 2-aminoquinoline-3-carboxylic acid derivatives and the data related to their in vitro screening.
Table 2. Structure of 2-oxo-1,2-dihydroquinoline-3-carboxylic acid derivatives and the data related to their in vitro screening.
Table 3. Structure of tetrazolo[1,5-a]quinoline-4-carboxylic acid derivatives and the data related to their in vitro screening.
Figure 2. Compound 5k (violet) in ATP-binding site of CK2 (the complex was obtained with molecular docking, hydrogen bonds are indicated by green dotted lines).
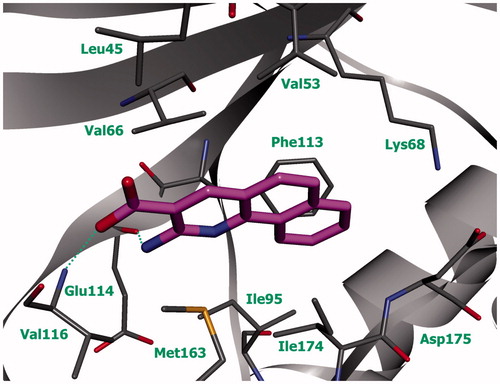
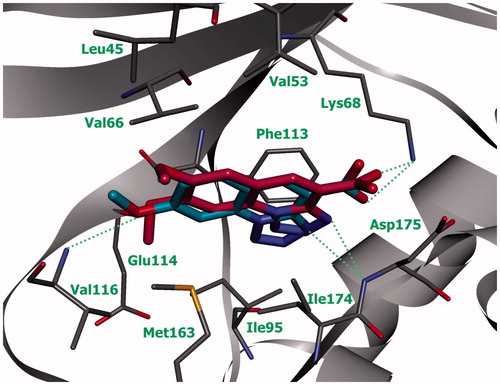
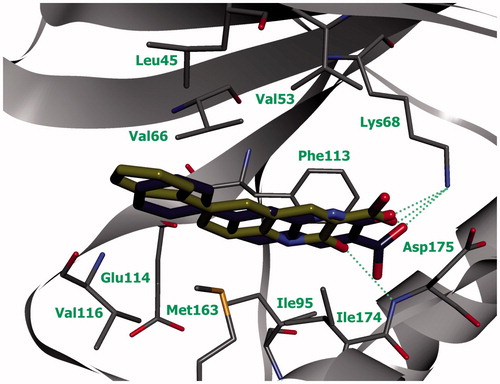
Figure 3. Compounds 6а (green) and 6b (blue) in ATP-binding site of CK2 (the complex was obtained by molecular docking, hydrogen bonds are indicated by green dotted lines).