Figures & data
Table 1. Analytical data of derivatives EMAC 3039–3064.
Table 2. 1H NMR data of derivatives EMAC 3039–3064.
Table 3. Activity of compounds EMAC 3039–3063 on HIV-1 RT-associated enzymatic functions RDDP and RNase H.
Figure 1. Synthetic pathway to compounds EMAC 3039–3063; reagents and conditions: (i) methylisothiocyanate, hydrazine hydrate, ethanol, rt; (ii) 1-amino-3-methylisothiourea, substituted isatin, ethanol, reflux; (iii) 2, substituted acetophenones, isopropanol, rt.
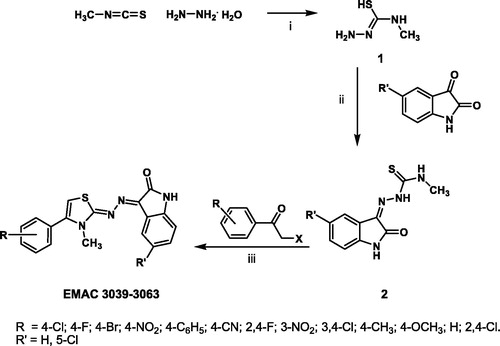
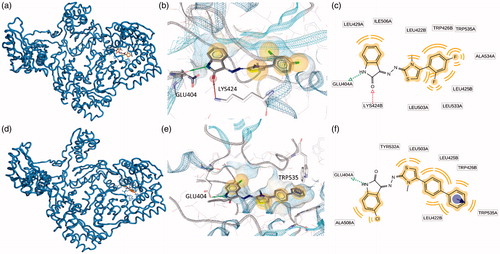
Figure 2. Putative binding modes of EMAC2045 and EMAC2056 and critical residues individuated for their binding in the pocket 1: (a, d) EMAC2045-HIV-1 RT complex and EMAC2056-HIV-1 RT complex; (b and e) close-up into the EMAC2045 and EMAC2056 binding site; (c and f) 2D depiction of EMAC2045 and its respective interactions with RT residues. pale yellow sphere indicates hydrophobic interactions with lipophilic residues. Red arrow indicates a hydrogen bond (HB) acceptor interaction, while the violet sphere represents the aromatic π − π stacking interaction.
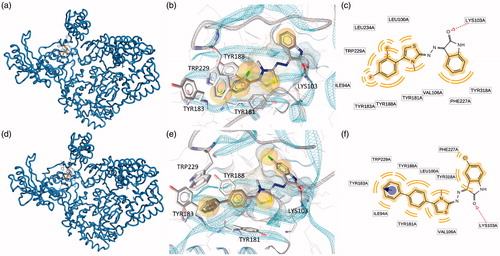
Figure 3. Putative binding modes of EMAC2045 and EMAC2056 and critical residues individuated for their binding in the pocket 2: (a, d) EMAC2045-HIV-1 RT complex and EMAC2056-HIV-1 RT complex; (b and e) close-up into the EMAC2045 and EMAC2056 binding site; (c and f) 2D depiction of EMAC2045 and its respective interactions with RT residues. pale yellow sphere indicates hydrophobic interactions with lipophilic residues. Red arrow indicates a hydrogen bond (HB) acceptor interaction, green HB donor, while the violet sphere represents the aromatic π − π stacking interaction.