Figures & data
Figure 1. Boxplots of some selected statistically significant (p > .05) metabolites from chamomile infusion indicating normalised intensity differences before (red) and after (green) enzymatic treatment. Esculin (1.33_340.0800n), luteoloside (2.44_448.1009n), rutin (2.14_611.1610m/z), esculetin (1.71_179.0343m/z), luteolin (2.98_287.0552m/z) and umbelliferone (0.60_163.0394m/z). All features were observed in the positive ionisation mode.
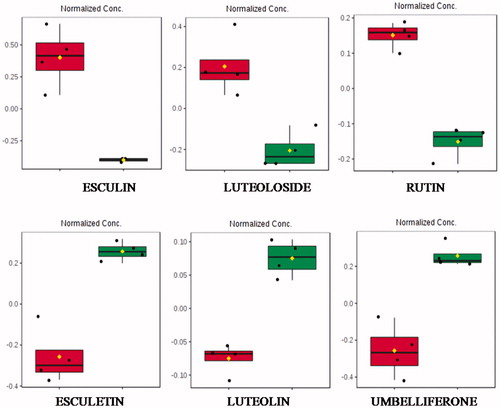
Table 1. Putative identification of statistically significant secondary metabolites from chamomile infusion and the relative tendency after the enzymatic treatment.
Table 2. DPPH• scavenging activity (%) and kinetic parameters of inhibitory effect of chamomile infusion toward digestive enzymes activities in the absence (control) and presence of the inhibitors (40 and 60 µM).
Figure 2. Lineweaver–Burk plot of glucosidase (A) and pancreatic lipase (B) activities. V is initial velocity and [S] is the concentration of substrate. The values were shown in absence (■) and presence of enzymatically modified chamomile infusion at 40 µM (●) and 60 µM (▲).The values are means of triplicate determinations, and the error bars indicate SD (n = 3).
![Figure 2. Lineweaver–Burk plot of glucosidase (A) and pancreatic lipase (B) activities. V is initial velocity and [S] is the concentration of substrate. The values were shown in absence (■) and presence of enzymatically modified chamomile infusion at 40 µM (●) and 60 µM (▲).The values are means of triplicate determinations, and the error bars indicate SD (n = 3).](/cms/asset/fafe261f-c185-43c7-9bdf-0c9c885f046c/ienz_a_1681989_f0002_b.jpg)
