Figures & data
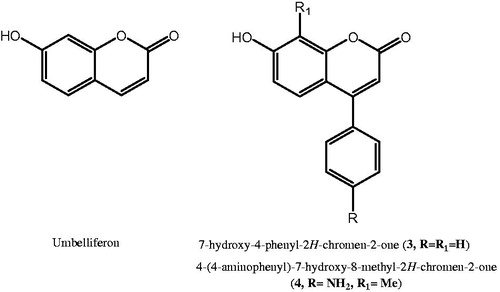
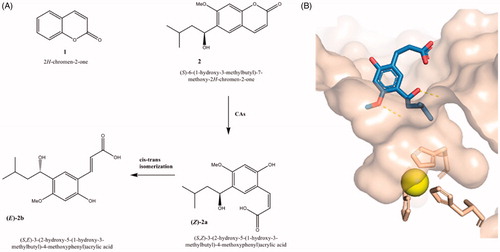
Figure 1. (A) Chemical structures of coumarin 1, 6-(1-S-hydroxy-3methylbutyl)-7-methoxy-2H-chromen-2-one (2) and corresponding hydrolytic products 2a and 2b; (B) cocrystal structure of hydrolytic product of 6-(1-S-hydroxy-3methylbutyl)-7-methoxy-2H-chromen-2-one (2) in complex with hCA II (PDB code: 3F8E)Citation12.
Scheme 1. (i) H2SO4, 0 °C to rt, 18–22 h; (ii) A: (MeCO)2O, H2SO4, Et3N, 0 °C to rt, 2 h; B: PheCOCl or EtCOCl, Et3N, DCM, rt, 24 h; (iii) AlCl3, 320 °C, 6 h; (iv) NH2NH2-H2O, Pd/C, EtOH, 70 °C, 1 h.
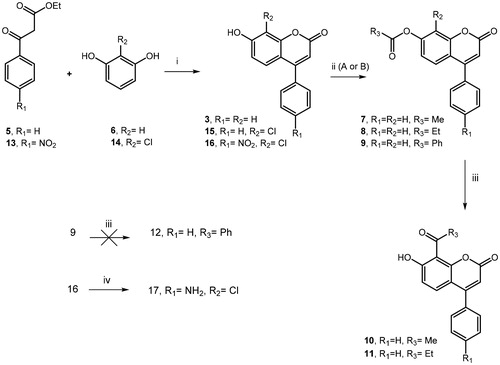
Table 1. Inhibition data for hCA I, II, IX and XII with compounds 7–11, 15 and 17 as well as umbelliferon and 3–4 as reference compoundsa.
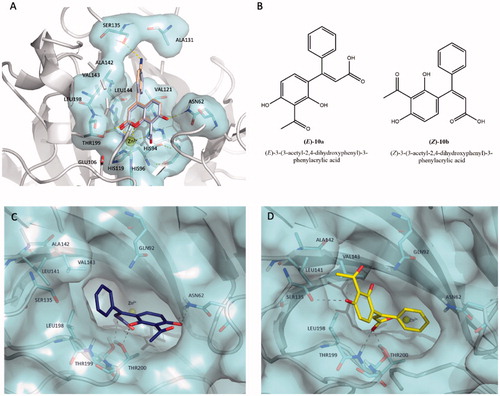
Figure 3. (A) Predicted binding mode of compound 10 (coloured in light blue) overlaid with compound 17 (coloured in wheat) into hCA XII cleft (PDB code 1JCZ).Citation42 (B) Chemical structures of hydrolytic forms 10a and 10b. Predicted binding mode of (E)-10a (C, coloured in blue) and (Z)-10b (D, coloured in yellow) as hydrolysed forms of coumarin 10 into hCA XII cleft (PDB Code: 1JCZ)Citation42. Compounds and crucial residues are shown as sticks; dashed lines represent hydrogen bond interactions. The protein structure is shown as pale-cyan surface and light grey cartoons. Zinc ion is depicted as a yellow sphere. Figures made by Pymol (https://pymol.org).