Figures & data
Figure 1. Flow chart. Simplified sketch of the performed protocol. HDF-cells are cultivated in 25 cm2 flask; upon confluence transfer into six 6-multiwell plates (working volume: 1.90–2.90 mL, surface/well: 9.6 cm2); exposure of three plates to the uwf-EM signal (for 10 min, distance: approx. 5 mm from the log-antenna); incubation of all six plates in different compartments of the incubator for up to 96 hours; execution of PCR (determination of HDF-viability and genetic expression of IL-6, TNFa, MMP-1, collagen and elastin) followed by a repetition of the 10 min uwf-EM treatment cycle on exposed cells every 24 hours. Note: the HDF-well in this plate was used as for monitoring purposes only. The HDF-well in this plate was used as for monitoring purposes only.
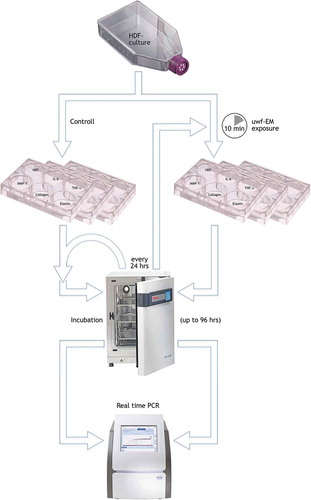
Table 1. Sense and antisense primers as obtained by real-time PCR.
Table 2. Two sided t- and p-values of t-test statistics. In accordance with the figures below, the diverging trends of controls and uwf-EMF-exposed cultures with a significance level of p < α (0.05) yields a non-significant difference in HDF data. However, H0 is rejected in the cases of IL-6, TNF-α and MM-1 (shaded areas); this is in line with tcrit being ≥2.777 among all data sets; this is further supported by tstat > tcrit leading also to the rejection of H0.
Figure 2. Cell viability measurement. HDF viability after treatment with and without uwf-EM exposure.
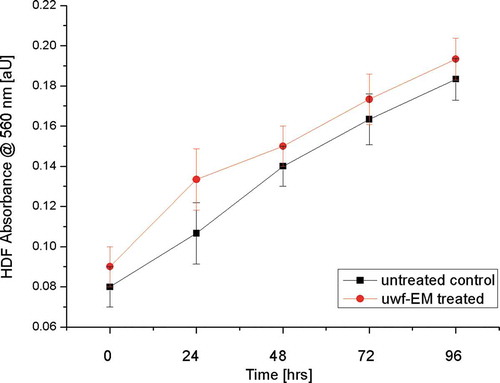
Figure 3. Real-time PCR analysis using specific primers for cytokines. Relative IL-6 gene expression from HDF (left) as well as TNF-α gene expression from HDF (right), treated with and without uwf-EM signals.
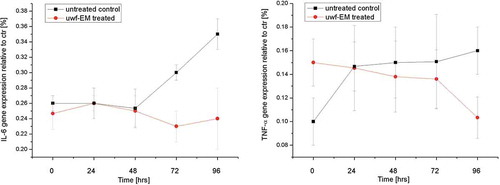
Figure 4. Real-time PCR analysis using specific primers for MMP-1, collagen type-I and elastin. Relative MMP-1 expression from HDF (top left), relative collagen type-1 gene expression from HDF (top right) and relative elastin gene expression from HDF (bottom), treated with and without uwf-EM signals.
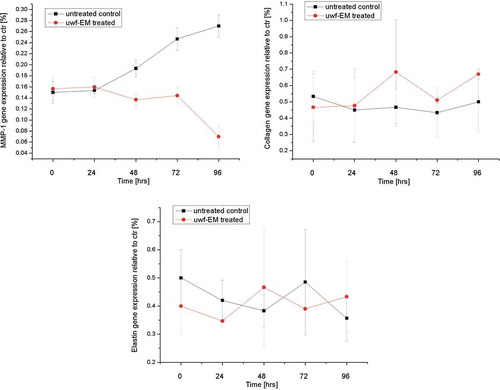
Table 3. Magnetic flux densities of background and FRACTOS logarithmic antenna in three repeats. Background values denote signal strength with full set-up yet at min. signal intensity, whereas 100% measurements denote maximum intensity of the emitted signal originating from the log-antenna.