Figures & data
Figure 1. Sensitivity of melanoma cell lines to anti-MEK inhibitors. (A) Short-term growth-inhibition assay of the indicated cell lines (SKMel10, M311, SKMel2, M376, IGRMEL1) treated with increasing concentrations of trametinib or cobimetinib. Cell viability was determined using the WST-1 cell proliferation assay. The data are presented as the mean +/− SEM (n = 3). (B) Long-term colony formation assay of the indicated cell lines. Cells were grown in the absence or presence of trametinib or cobimetinib at the indicated concentrations for 7–14 days. For each cell line, all dishes were fixed at the same time, stained and photographed.
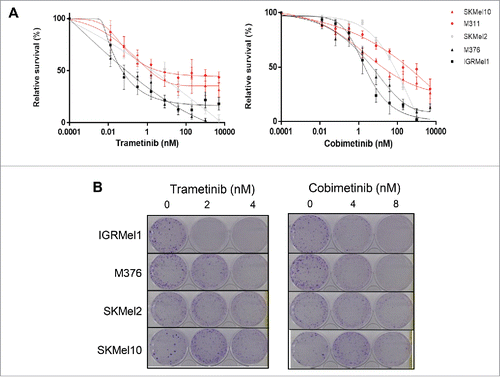
Figure 2. The formation of the eIF4F translation initiation complex is associated with resistance to MEK inhibitors. (A) eIF4E–eIF4G interactions detected by proximity ligation assay (PLA) in trametinib/cobimetinib-treated or untreated cell lines. The interactions were visualized as red spots. (B) PLA quantification showing the number of eIF4E-eIF4G interactions by cell. The data are presented as the mean s.d (n = 4), and differences were assessed with Student's t-test (* p < 0,05; ** p < 0,01; *** p < 0,001).
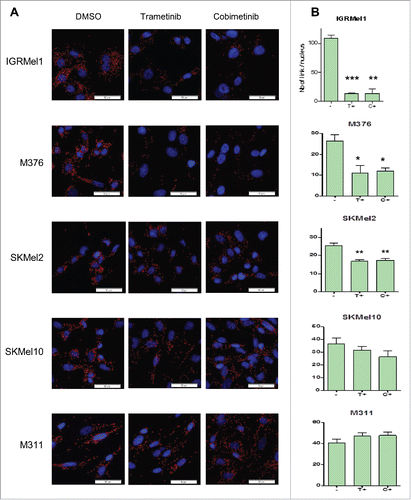
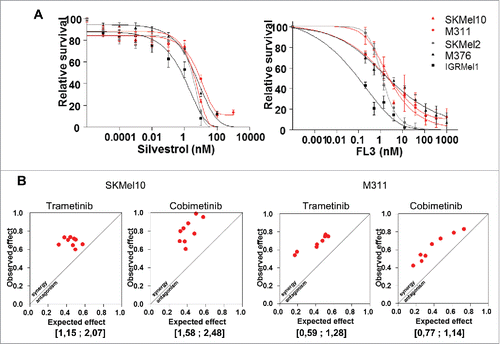
Figure 3. Flavaglines sensitize NRAS-mutated resistant cell lines and synergize with MEK inhibitors. (A) Short-term growth-inhibition assay of the indicated cell lines treated with increasing concentrations of silvestrol or FL3. Cell viability was determined using the WST-1 cell proliferation assay. The data are presented as the mean +/− SEM (n = 3). (B) Isobologram of the effect of the combination of trametinib or cobimetinib plus FL3 at fixed concentration (4,1 nM) on SKMel10 and M311 cell lines. The Bliss index is shown in square brackets.