Figures & data
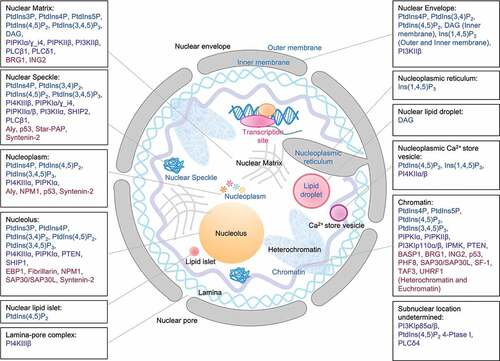
Table 1. Nuclear targeted PIs: types, locations, and response to cell stress
Table 2. Nuclear PI-metabolizing enzymes, their substrates, nuclear location and response to cell stress
Table 3. Nuclear PI-binding proteins, their interacting PIs, associated PI enzymes, nuclear location, response to cell stress and functions