Figures & data
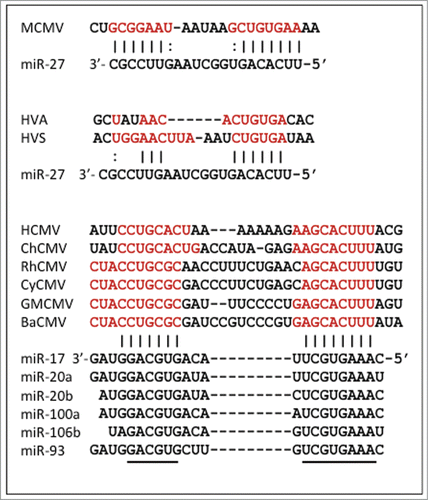
Figure 1. Phylogenetic relationships among representative mammalian herpesviruses, showing the branches on which 3 distinct miRNA regulators appear to have evolved (circles), and 4 occasions where IL10 genes have been acquired (triangles). Ancestral branches leading to the α, β and gamma subfamilies of herpesviruses are indicated. Virus acronyms are explained, and sequence sources are given, in Supplementary Table 1.
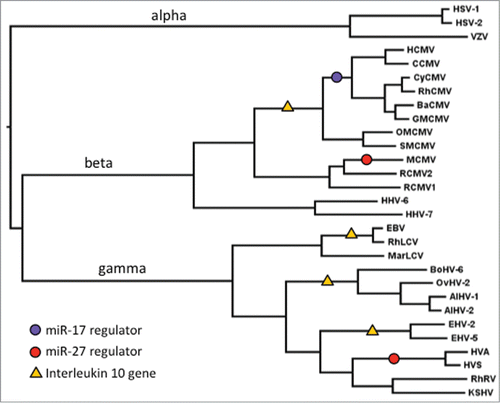
Figure 2. miRNA binding sites in herpesviruses. Sites within viral RNAs that are complementary to miRNAs are shown in red; black lines (or dots for G-U) show sites where potential binding is conserved. Conserved sequences in the miR-17 family members are denoted with a black line. Virus acronyms are explained, and sequence sources are given, in Supplementary Table 1.