Figures & data
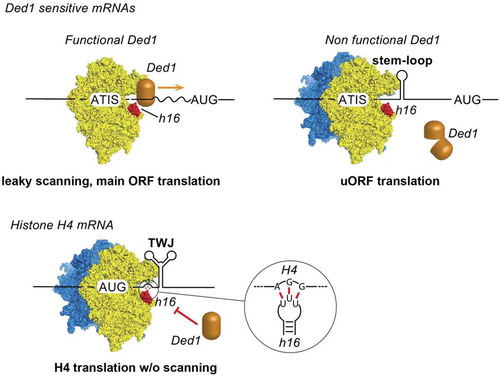
Figure 1. Examples of mRNAs that are translated by the START mechanism. In yeast, Ded1 sensitive mRNAs contain uORF that begin with an Alternative Initiation Site (ATIS). Secondary RNA structures downstream of ATIS stall the scanning complex and trigger translation initiation on ATIS [Citation5]. In metazoan histone H4 mRNA, a highly stable Three Way Junction (TWJ) positions the AUG start codon in the P site of the ribosome [Citation6,Citation10]. In ALS/FTD patients, C9ORF72 transcripts contained G4C2 repeats that fold into highly stable G-quartet structures. The G-quadruplexes stall the scanning complex and forces translation initiation on the near-cognate CUG codon [Citation7]. In some prokaryotic mRNAs, the ribosome is positioned on the AUG start codon by an upstream interaction between the 16S rRNA with the Shine-Dalgarno (SD) sequence and by a stable hairpin downstream of the AUG. Small regulatory RNAs regulate the folding of the downstream hairpins [Citation8]. During viral translation of EMCV genome, an RNA-protein complex formed by the viral protease 2A bound to a stem-loop structure guides the ribosome for −1 programmed frameshifting [Citation9].
![Figure 1. Examples of mRNAs that are translated by the START mechanism. In yeast, Ded1 sensitive mRNAs contain uORF that begin with an Alternative Initiation Site (ATIS). Secondary RNA structures downstream of ATIS stall the scanning complex and trigger translation initiation on ATIS [Citation5]. In metazoan histone H4 mRNA, a highly stable Three Way Junction (TWJ) positions the AUG start codon in the P site of the ribosome [Citation6,Citation10]. In ALS/FTD patients, C9ORF72 transcripts contained G4C2 repeats that fold into highly stable G-quartet structures. The G-quadruplexes stall the scanning complex and forces translation initiation on the near-cognate CUG codon [Citation7]. In some prokaryotic mRNAs, the ribosome is positioned on the AUG start codon by an upstream interaction between the 16S rRNA with the Shine-Dalgarno (SD) sequence and by a stable hairpin downstream of the AUG. Small regulatory RNAs regulate the folding of the downstream hairpins [Citation8]. During viral translation of EMCV genome, an RNA-protein complex formed by the viral protease 2A bound to a stem-loop structure guides the ribosome for −1 programmed frameshifting [Citation9].](/cms/asset/3f18cc09-1de2-4224-8ff4-b62bd8ad6735/krnb_a_1518855_f0001_oc.jpg)
Figure 2. Ribosomal helix h16 from 18S rRNA interacts with Ded1 helicase. Ded1 is located in the scanning complex by interacting with helix h16. Ded1 unwinds secondary structures in the 5ʹ UTR during 5ʹ-3ʹ scanning until the AUG start codon is found. When Ded1 is non-functional, secondary structures in the 5ʹUTR promote translation initiation on ATIS and lead to uORF translation and down-regulation of the main ORF. During histone H4 mRNA translation, an AGG triplet upstream of the Three-Way Junction (TWJ) interacts specifically with the loop of helix h16 and thereby prevents Ded1 interaction . The ribosome is positioned directly on the AUG start codon with the help of TWJ and without scanning.