Figures & data
Figure 1. Two-step degradation reaction of pseudouridine. PUKI and PUMY are responsible for pseudouridine catabolism. Atomic numbering of the nucleobase is indicated.
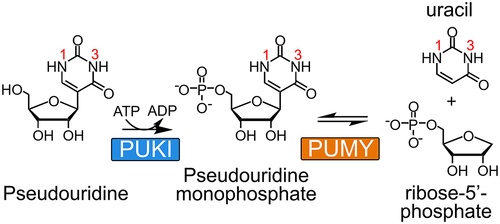
Table 1. Data collection and refinement statistics.
Figure 2. Structural features of the AtPUMY holoenzyme.
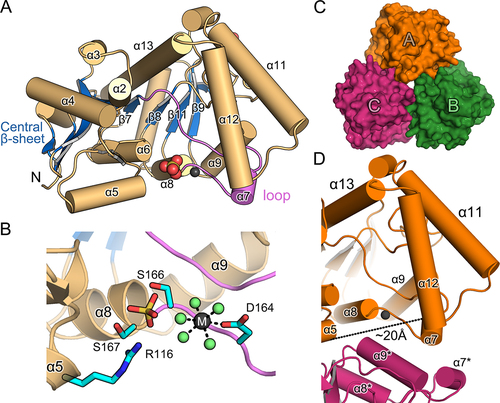
Figure 3. Ligand-binding modes and conformational changes in AtPUMY.
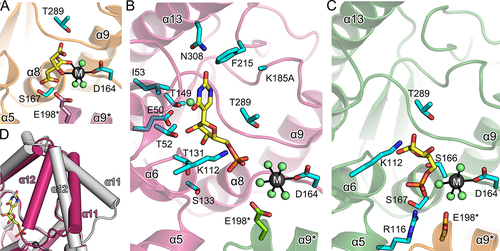
Figure 4. Enzymatic properties of AtPUMY.
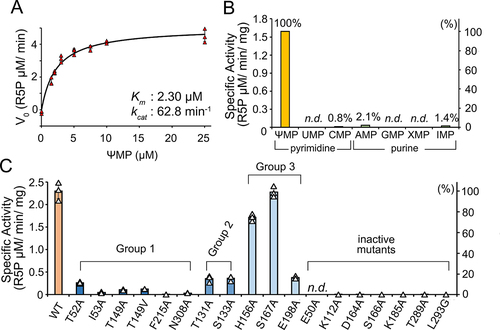
Table 2. Kinetic parameters of AtPUMY mutants.
Supplemental Material
Download MS Word (1.5 MB)RHEE_8K06.pdf
Download PDF (3.2 MB)RHEE_8K05.pdf
Download PDF (1.2 MB)RHEE_8K07.pdf
Download PDF (2.7 MB)Data availability statement
The atomic coordinates and structural factors have been deposited in the Protein Data Bank (http://www.rcsb.org) under ID code 8K05, 8K06, and 8K07.
