Figures & data
Figure 1. Atg11 is required for glucose starvation-induced autophagy. GFP-Atg8 was expressed in the yeast strains listed from (A–D). Cells were grown to the log-growth phase, then subjected to nitrogen starvation or glucose starvation for 4 h. Autophagic activity was assessed by western blot analysis of GFP-Atg8 cleavage
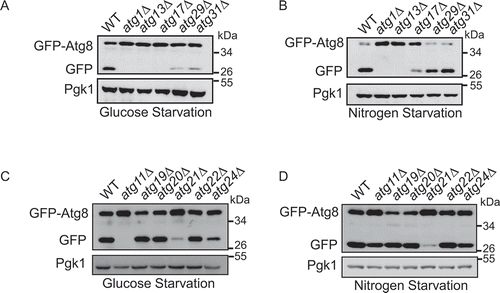
Figure 2. Atg11 is essential for activating Atg1 in glucose starvation conditions. (A) Atg1-3XFLAG was expressed in wild type, atg11∆, atg13∆, atg17∆ yeast strains. Cells were subjected to nitrogen starvation (SD-N) or glucose starvation medium (SD-G) for 1 h. Cell lysates were immunoprecipitated with anti-FLAG magnetic beads. An In vitro phosphorylation assay was carried out with MBP protein as substrate and immunoprecipitated-Atg1 as kinase for analyzing Atg1 kinase activity. S.E and L.E stands for short exposure and long exposure, respectively. (B) Cells expressing HA-Snf1 were cultured in full medium, SD-N, SD-G, or treated with rapamycin for 1 h. Cell lysates were then immunoblotted with anti-p-Thr172 PRKAA/AMPKα antibody or anti-HA antibody. (C) His6-tagged trigger factor (TF)-Atg1 and TF-Mec1-1300-2000 aa protein was purified with a Ni2+ column from E. Coli; Snf1-WT and Snf1-KD protein were purified with TAP tag from glucose starvation-treated snf1∆ yeast cells expressing Snf1-3XFLAG-TEV-ZZ or Snf1-KD-3XFLAG-TEV-ZZ. Purified Snf1-WT or Snf1-KD were incubated with TF-Atg1 or TF-Mec1-1300-2000aa protein and ATP-γ-S in Snf1 kinase buffer at 30°C for 30 min, and followed by PNBM treatment for another hour. The phosphorylation of Atg1 was detected using an anti-thiophosphate ester (anti-thioP) antibody. Snf1 kinase activity was monitored by immunoblotting with an anti-p-PRKAA/AMPKα (Thr172) antibody. (D) The Atg1-3XFLAG protein was purified using anti-FLAG magnetic beads from BY4741 and snf1∆ yeast strain under full medium and SD-G. An in vitro Atg1 phosphorylation assay was carried out with MBP protein as a substrate and immunoprecipitated-Atg1 as a kinase. (E) HA-Snf1 and Atg1-3XFLAG were co-expressed in wild type and atg11∆ yeast strains. Cells were grown to the log-growth phase and then subjected to glucose starvation for 1 h. Cell lysates were immunoprecipitated with anti-FLAG magnetic beads and analyzed by western blot with the indicated antibodies. IC: mouse IgG1 isotype control
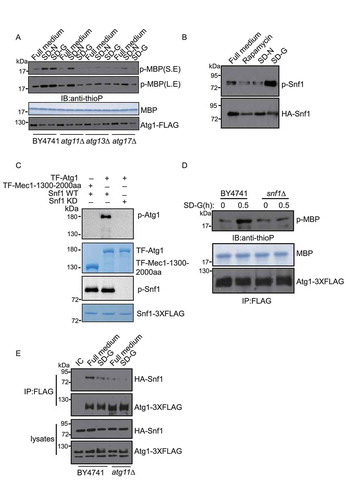
Figure 3. Atg11 is required for the formation of the Atg17-Atg31-Atg29 complex in glucose starvation conditions. (A-C) Atg17-2XGFP, Atg29-2XGFP and Atg31-2XGFP were expressed in the wild type and atg11∆ strains. Cells were cultured in full medium, SD-N or SD-G for 1 h, then analyzed by microscopy. Scale bar, 2 µm. (D) Atg17-6XHA, Atg29-TAP and Atg31-2XGFP were co-expressed in wild type and atg11∆ yeast strains. Cells were grown to the log-growth phase and then subjected to SD-N or SD-G for 1 h. Cell lysates were immunoprecipitated with anti-HA magnetic beads and then analyzed with the indicated antibodies. IC: mouse IgG1 isotype control. (E) Cell co-expressing Atg31-2XGFP and Atg29-TAP in the wild type or atg11∆ strains were then subjected to SD-N and SD-G for 1 h. Cell lysates were immunoprecipitated with rabbit IgG agarose beads and then analyzed by western blot using the indicated antibody. (F) Gel filtration chromatography analysis by Superose 6 10/300 GL column for proteins lysed from wild type (WT) and atg11∆ yeast after cells were deprived from glucose for 1 h. Each fraction was detected by immunoblotting using anti-HA, anti-GFP and anti-Protein A antibodies. Positions of molecular weight (MW) between fractions are shown
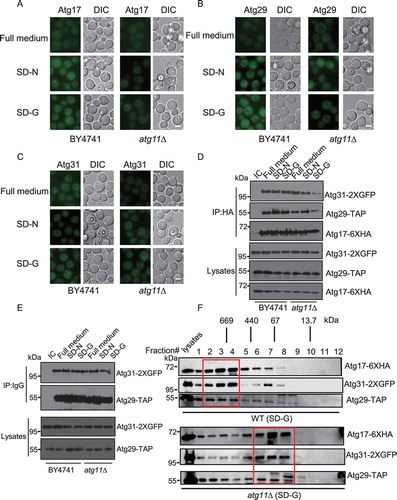
Figure 4. The recruitment of Atg9 vesicles to the PAS by Atg11 is required for glucose starvation-induced autophagy. (A) The scheme of the Atg11 domains and the deletions. CC1∆ involves the deletion of amino acids at the 272–321 position of the Atg11 protein, CC2∆ involves the deletion of amino acids at the 536–576 position, CC3∆ involves the deletion of amino acids at the 627–859 position, CC4∆ involves the deletion of amino acids at the 859–1178 position. Modified from [Citation18]. (B) Yeast two-hybrid analysis of the Atg11 and Atg9 interaction. The yeast strain AH109 was transformed with plasmids expressing AD-fused with Atg11, Atg11 CC2∆, Atg11 566-576aa∆, or Atg11I569E and plasmids expressing a BD-fused with the N terminus of Atg9. These strains were grown on SD-Leu-Trp (+Ade) and SD-Leu-Trp-Ade (−Ade) agar plates at 30°C for 3 d. (C) Cells co-expressing an empty vector, HA-Atg11 WT, HA-Atg11-CC2∆, or HA-Atg11I569E with Atg9-TAP in the atg11∆ strain were starved with glucose for 0 h and 1 h. Cell lysates were immunoprecipitated with rabbit IgG agarose beads and then analyzed by western blot using the anti-HA antibody. (D) Images of cells expressing Atg9-2XGFP and RFP-Ape1, which were starved in SD-G for 1 h. Scale bar, 2 µm. (E) Cells from (D) were quantified for the number of cells with Atg9-2XGFP dots co-localized with RFP-Ape1 dots. n = 300 cells pooled from three independent experiments. Data are presented as means ± SD. (F) Cells expressing GFP-Atg8 and Vph1-Cherry in atg11∆, HA-Atg11-WT, HA-Atg11-CC2∆, and HA-Atg11I569E strains were cultured in SD-N or SD-G medium for 4 h. Autophagic activity was assessed by the quantification of the translocation of GFP-Atg8 into vacuoles. Scale bar, 2 µm. (G) Cells from (F) were analyzed for the translocation of GFP-Atg8 into vacuoles. n = 300 cells pooled from three independent experiments. Data are presented as means ± SD. (H) Cells from (F) were analyzed by western blot for GFP-Atg8 cleavage
![Figure 4. The recruitment of Atg9 vesicles to the PAS by Atg11 is required for glucose starvation-induced autophagy. (A) The scheme of the Atg11 domains and the deletions. CC1∆ involves the deletion of amino acids at the 272–321 position of the Atg11 protein, CC2∆ involves the deletion of amino acids at the 536–576 position, CC3∆ involves the deletion of amino acids at the 627–859 position, CC4∆ involves the deletion of amino acids at the 859–1178 position. Modified from [Citation18]. (B) Yeast two-hybrid analysis of the Atg11 and Atg9 interaction. The yeast strain AH109 was transformed with plasmids expressing AD-fused with Atg11, Atg11 CC2∆, Atg11 566-576aa∆, or Atg11I569E and plasmids expressing a BD-fused with the N terminus of Atg9. These strains were grown on SD-Leu-Trp (+Ade) and SD-Leu-Trp-Ade (−Ade) agar plates at 30°C for 3 d. (C) Cells co-expressing an empty vector, HA-Atg11 WT, HA-Atg11-CC2∆, or HA-Atg11I569E with Atg9-TAP in the atg11∆ strain were starved with glucose for 0 h and 1 h. Cell lysates were immunoprecipitated with rabbit IgG agarose beads and then analyzed by western blot using the anti-HA antibody. (D) Images of cells expressing Atg9-2XGFP and RFP-Ape1, which were starved in SD-G for 1 h. Scale bar, 2 µm. (E) Cells from (D) were quantified for the number of cells with Atg9-2XGFP dots co-localized with RFP-Ape1 dots. n = 300 cells pooled from three independent experiments. Data are presented as means ± SD. (F) Cells expressing GFP-Atg8 and Vph1-Cherry in atg11∆, HA-Atg11-WT, HA-Atg11-CC2∆, and HA-Atg11I569E strains were cultured in SD-N or SD-G medium for 4 h. Autophagic activity was assessed by the quantification of the translocation of GFP-Atg8 into vacuoles. Scale bar, 2 µm. (G) Cells from (F) were analyzed for the translocation of GFP-Atg8 into vacuoles. n = 300 cells pooled from three independent experiments. Data are presented as means ± SD. (H) Cells from (F) were analyzed by western blot for GFP-Atg8 cleavage](/cms/asset/68577456-fe80-42d7-a81c-dde261227b99/kaup_a_1719724_f0004_oc.jpg)
Figure 5. The CC1, CC2, CC3, and CC4 domains of Atg11 are required for glucose starvation-induced autophagy. (A) Cells expressing GFP-Atg8 and Vph1-Cherry in atg11∆, HA-Atg11 WT, HA-Atg11-CC1∆, HA-Atg11-CC2∆, HA-Atg11-CC3∆, and HA-Atg11-CC4∆ strains were cultured in SD-N or SD-G medium for 4 h. Autophagic activity was assessed for the translocation of GFP-Atg8 into vacuoles. n = 300 cells pooled from three independent experiments. Data are presented as means ± SD. (B) Autophagic activity from (A) was assessed using western blot analysis of GFP-Atg8 and Ape1 cleavage. (C-E) Cells co-expressing an empty vector, HA-Atg11 WT, HA-Atg11-CC1∆, HA-Atg11-CC2∆, HA-Atg11-CC3∆, or HA-Atg11-CC4∆ with Atg1-3XFLAG, Atg29-3XFLAG, or Atg31-3XGLAG in the atg11∆ strain were grown to the log phase and then subjected to glucose starvation for 1 h. Cell lysates were immunoprecipitated with anti-FLAG magnetic beads and then analyzed by western blot using the indicated antibody. (F) Cells co-expressing an empty vector, HA-Atg11 WT, HA-Atg11-CC1∆, HA-Atg11-CC2∆, HA-Atg11-CC3∆, or HA-Atg11-CC4∆ with Snf1-3XFLAG in the atg11∆ strain were grown to the log phase. Cell lysates were immunoprecipitated with anti-FLAG magnetic beads and then analyzed by western blot using the indicated antibody
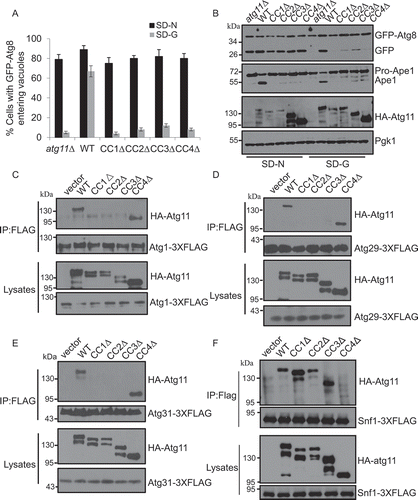
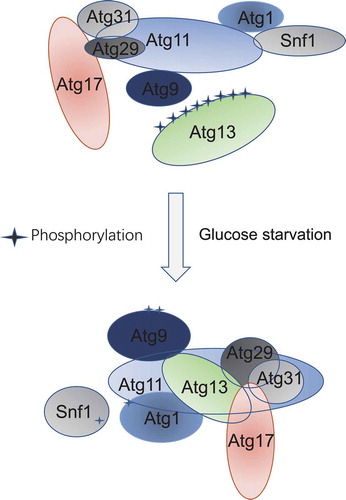
Figure 6. Model of Atg11 participating in initiation of glucose starvation-induced autophagy. In response to glucose starvation, Atg11 participates in the assembly of the PAS by regulating the association of Atg17 with Atg29-Atg31. Simultaneously, Atg1 is activated by Atg11 mediating binding of Atg1 with Snf1, and Atg9 vesicles are recruited to the PAS by Atg11 to initiate glucose starvation-induced autophagy