Figures & data
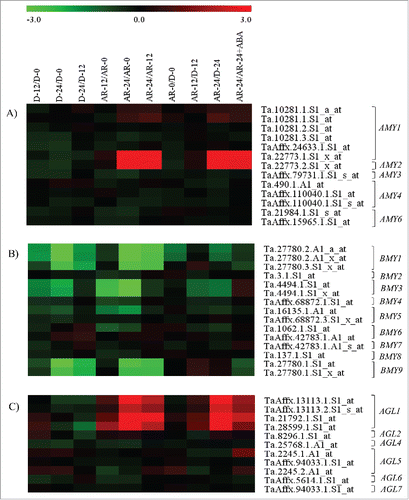
Figure 1. Heat maps showing changes in the expression of probesets annotated as AMY (A), BAM (B) and AGL (C) genes. Log2 scaled fold changes in the expression of probesets during imbibition of dormant (D-12/D-0, D-24/D-0 and D-24/D-12) and after-ripened seeds (AR-12/AR-0, AR-24/AR-0, AR-24/AR-12), between dormant and after-ripened seeds in both dry and imbibed states (AR-0/D-0, AR-12/D-12 and AR-24/D-24) and between water and ABA imbibed after-ripened seeds (AR-24/AR-24+ABA). Expression values in log2 fold changes are represented by the negative and positive numbers on the bar and the color scale at the top of each heat map; higher and lower expression levels of respective probesets are represented by red and green colors, respectively. Log2 and linear scaled fold changes in expression of the probesets and the respective P values can be found in Table S1.