Figures & data
Table 1. Biomarker classification [Citation71] and approaches to assess quantitative information.
![Figure 1. The combinatory mapping approach. (Redrawn from [Citation23] with permission of Springer).](/cms/asset/76a631c9-1d13-46ab-b77d-d44a73d5065a/iedc_a_1380623_f0001_oc.jpg)
![Figure 2. The structure of the generic CNS PBPK model Structure: Black: Plasma PK model, Red: CNS PBPK model. Parameters: Black: estimated plasma PK parameters; Blue: system-specific parameters; Green: drug-specific parameters; Purple: combination of system-specific and drug-specific specific parameters. (Redrawn from [Citation39] with permission of Wiley & Sons).](/cms/asset/54b728fa-5f77-400d-94e7-5b81cb78168e/iedc_a_1380623_f0002_oc.jpg)
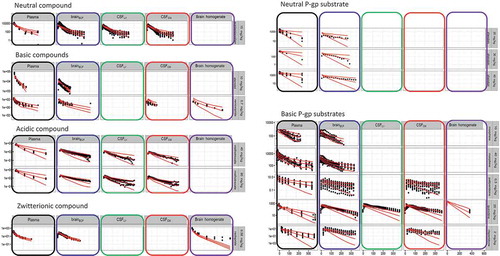
![Figure 5. Compared plasma pharmacokinetics of high affinity compounds (top row) and their low affinity analogues (bottom row). The circles represent observed plasma concentrations in rats, the lines represent model predictions. (Reprinted from [Citation45] with permission of ASPET).](/cms/asset/07e7759f-ff48-48c8-8bca-24a066bccd4b/iedc_a_1380623_f0005_b.gif)
