Figures & data
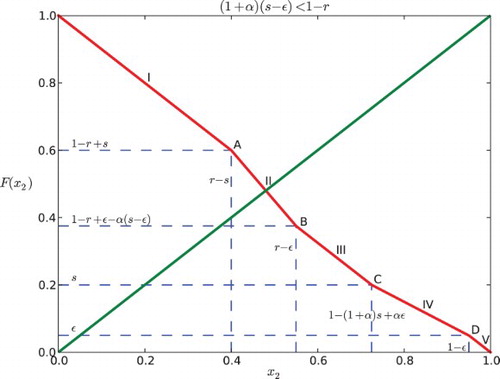
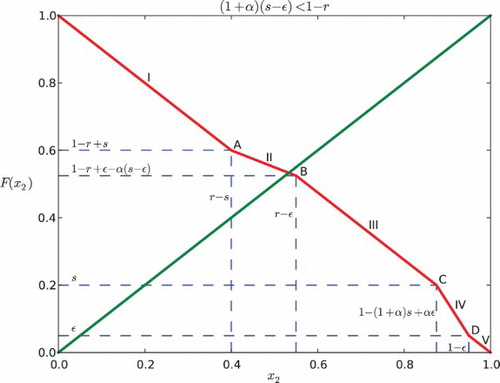
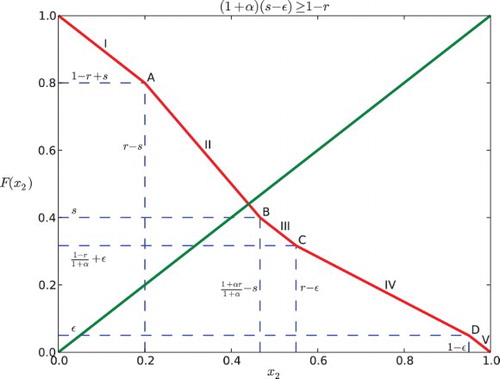
Fig. 6. Regions of parameter space for the k cyclic solutions with k=M+1. Here the feedback is negative with β=−.2 and the gap is ε=0.03. The widest diagonal band contains the parameters for k=2 and is partitioned into six cases. Note that case VI disobeys the constraint ε<r−s. The rest of the diagonal bands contain parameters for k=3, 4, … and each of them is partitioned into five cases.
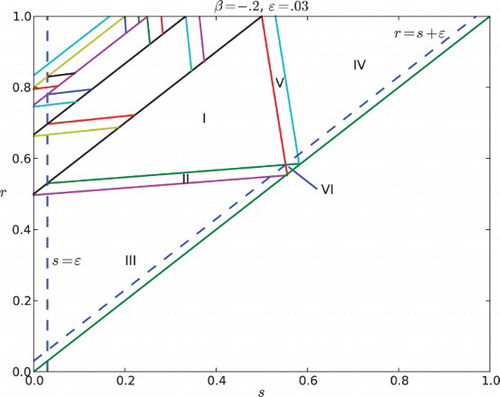
Fig. 7. The number of clusters realized in a cell cycle simulation (a) with no gap and (b) with a gap of ε=0.05 when s≥0.01 and ε=0.5 s when s<0.1. In both plots, the feedback is linear and negative with f(I)=−I. Note the strong agreement between the two plots.
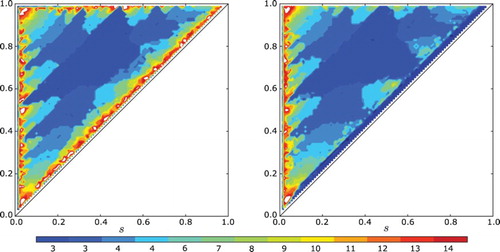
Fig. 8. Clustering of n=120 equi-distributed cells.(a) For s=0.2, r=0.6 and ε=0.03, the cells converge to two clusters. (b) For s=0.2, r=0.8 and ε=0.01, the cells converge to three clusters. For all simulations the feedback was taken to be f(I)=−I. Note in (a) that they first appear to form four clusters, but merge into two clusters.
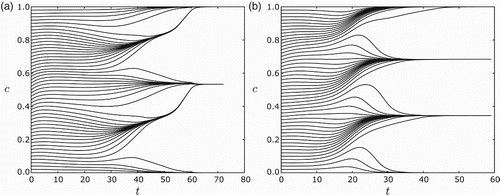
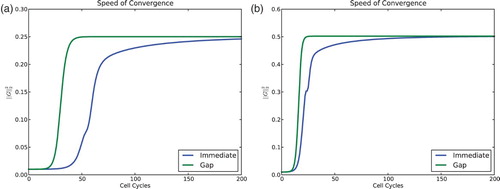
Fig. 9. Comparison of convergence speed for the model with and without a gap of ε=0.01 and n=60 cells. Norm squared of the gap vector G vs. time. (a) For s=0.2 and r=0.9 both models converge to four clusters. (b) For s=0.3 and r=0.7 both models converge to two clusters. For all simulations the feedback was taken to be f(I)=−I, and the initial condition was evenly distributed – the global minimum for 0. The model with gaps converges to the clustered solutions much more quickly.

![Fig. 10. The final configuration of cells in systems with gap sizes varying in [0.1, 4.9].](/cms/asset/2f70738f-fa44-4e6c-9920-fac61f713f57/tjbd_a_904526_f0010_b.gif)