Figures & data
FIGURE 1. (A) Structures of bi-, tri-, and tetraantennary N-linked glycans found in PrPC and PrPSc that could be bisected (b) or nonbisected (n).Citation18,20,21 Facultative fucosialtion and sialylation are shown within parenthesis with several sialic acid residues per glycan indicated. (B) Structures of pentatantennary and nonconventional N-liked glycans that can accommodate up to 5 sialic acid residues.Citation28
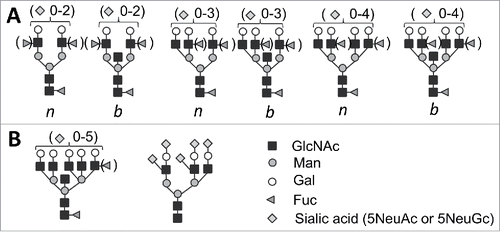
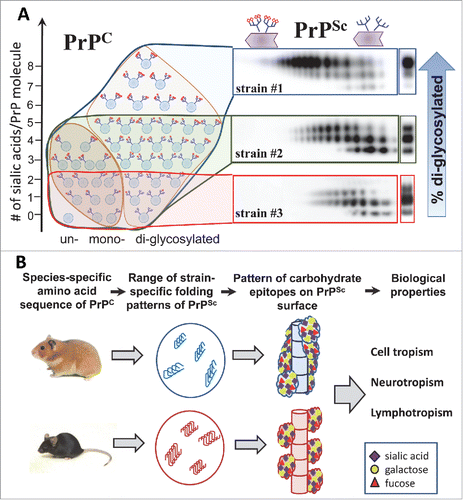
FIGURE 2. (A) Schematic diagram illustrating that PrPSc strains recruit PrPC isoforms selectively according to PrPC glycosylation and sialylation status (adopted fromCitation12). Strain #1 recruits sialoglycoforms of PrPC without noticeable preferences. Hypersialylated and diglycosylated PrPC are preferentially excluded from the strain #2 and even more so from the strain #3. As a result, the proportion of hyper- versus hyposialylated molecules within PrPSc (illustrated by the 2D Western blots), as well as the ratios of di-, mono- and unglycosylated glycoforms (shown on 1D Western blots on right hand side), changes in a strain-specific manner. PrPC molecules are shown as blue circles and sialic acid residues - as red diamonds. (B) Schematic diagram illustrates that species-specific amino acid sequences of PrPC determine the range of strain-specific folding patterns, which define the strain-specific patterns of functional carbohydrate epitopes on PrPSc surfaces within a particular host. Patterns of functional carbohydrate epitopes determine strain-specific biologic properties. PrPSc folding patterns are expected to be quite different for 2 groups of strains: (i) a group that can accommodate diglycosylated glycoforms and (ii) another group that selectively excludes diglycosylated glycoforms.