Figures & data
Table 1. Formulation and fluidity of liposomes mimicking normal (SCLL) and glycated (G-SCLL) epidermal lipid composition. Results are expressed as the mean ± SD of n = 3 replicates.* p < 0.05.
Figure 1. General properties of the glycated reconstructed epidermal model. Glycation was induced by hydration with various concentrations of GO from the basal side for 72 h. Viability of glycated 6D RHE was measured by alamar Blue® assay(a). ΔTEWL of glycated 6D RHE was measured using vaposcan (b). All results are expressed as the mean ± SD of n = 3 or n = 4 replicates, respectively. *p < 0.05, ***p < 0.001.
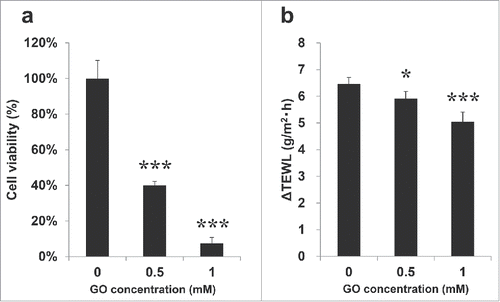
Figure 2. Changes in the content of epidermal lipids in glycated reconstructed epidermal model. Lipids contents of 6D RHE were determined after 72 h exposure to glyoxal by HPTLC: CER[NS] (a), CER[NP] (b), CER[AS] (c), CER[AP]a (d), CER[AP]b (e), cholesterol (f), and fatty acids (g). All results are expressed as the mean ± SD of n = 3 replicates. *p < 0.05, **p < 0.01, ***p < 0.001.
![Figure 2. Changes in the content of epidermal lipids in glycated reconstructed epidermal model. Lipids contents of 6D RHE were determined after 72 h exposure to glyoxal by HPTLC: CER[NS] (a), CER[NP] (b), CER[AS] (c), CER[AP]a (d), CER[AP]b (e), cholesterol (f), and fatty acids (g). All results are expressed as the mean ± SD of n = 3 replicates. *p < 0.05, **p < 0.01, ***p < 0.001.](/cms/asset/0db93485-a1b4-4cff-b78b-f2d37ca742b8/kder_a_1338992_f0002_b.gif)
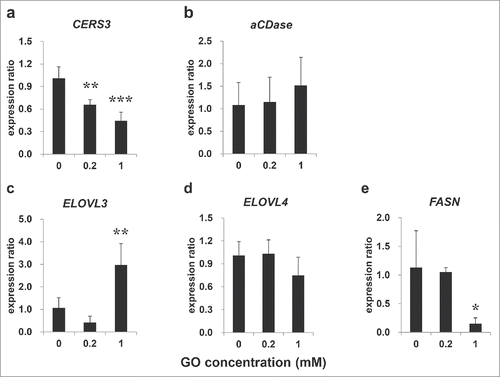
Figure 3. Expression of ceramide and fatty acid metabolism-associated genes. Expression of CERS3 (a), aCDase (b), ELOVL3 (c) ELOVL4 (d) and FASN (e) in EPISKIN after 72 h exposure to glyoxal as determined by real-time RT PCR. All results are expressed as the mean ± SD of n = 3 replicates. *p < 0.05, **p < 0.01, ***p < 0.001.