Figures & data
Figure 1. Malnourished media composition and study design. To generate malnourished organoids, certain media additives were eliminated to reduce nutritional content while maintaining cellular viability. Following establishment of media formulations to reduce protein, amino acid, and carbohydrate concentrations, both organoids and organoid-derived monolayers were incubated in the various media compositions and subsequently analyzed for the effects of malnourishment.
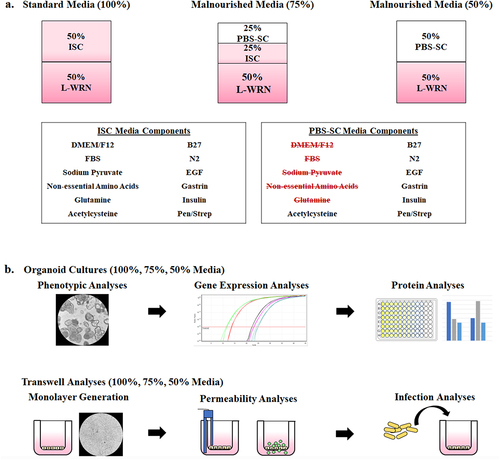
Figure 2. Phenotypic analysis of malnourished organoids.
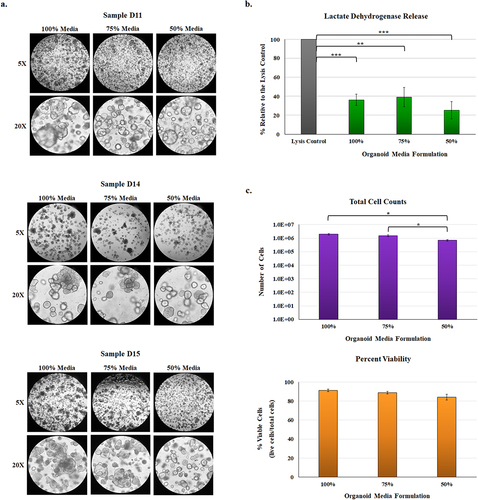
Figure 3. Gene expression analyses of malnourished organoids and patient biopsy samples with different BMIs. Quantitative RT-PCR was performed for the indicated genes. Data bars represent the average fold change ± the SEM. All data were normalized to the expression of the 18S housekeeping gene for each experiment. Statistical significance was determined with the Student’s T-test of the ΔCT and the treatment comparisons that are statistically different are indicated for each gene and comparison. Data were considered significant at a p value of < .05 (*, <.05; **, <.01; ***, <.001). Please note the different y-axes for each graph.
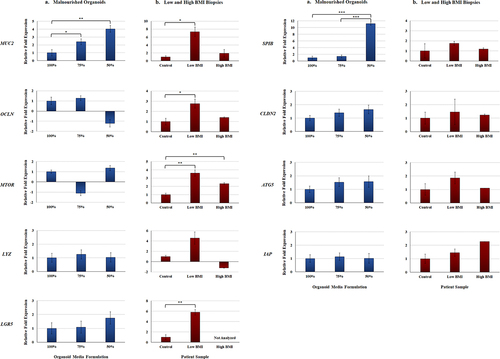
Figure 4. Protein expression in the malnourished organoids. Analyses were performed as a measure of protein expression in the organoids with the various media formulations.
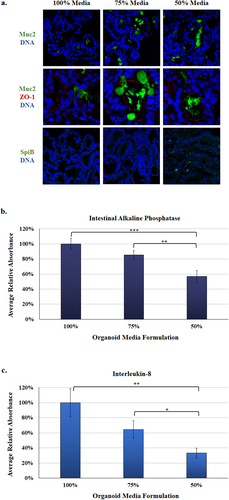
Figure 5. Malnourished monolayer permeability analysis. To measure integrity and permeability of the malnourished monolayers, mHIODEM were derived from the D11 and D14 organoid lines, and TEER and diffusion of FITC-dextran were measured. Each measurement occurred following monolayer differentiation with DAPT. Data represent three biologically independent experiments, each with at least two technical replicate wells. For both analyses, the average data ± SEM are plotted. Statistical significance was determined with the Student’s T-test comparing results from the 100% media to the 75% or 50% media. Data were considered significant at a p value of < .05 (*, <.05; **, <.01; ***, <.001). Please note the different y-axes for each graph.
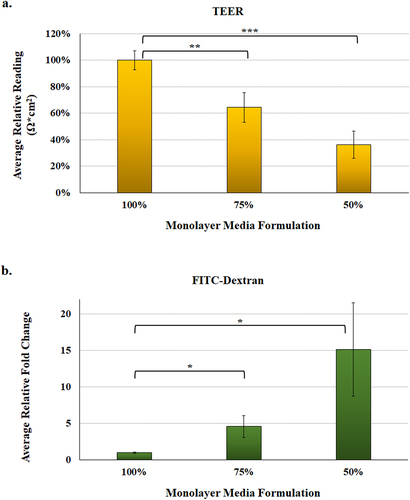
Figure 6. Malnourished monolayer infection analysis. Organoids were trypsinized, seeded onto transwells, and cultured in the 100%, 75%, and 50% media formulations. Following 11 days of culturing and a 48-hour differentiation treatment with DAPT, monolayers were subsequently infected with S. flexneri strain 2457T. Following apical administration of the bacteria, plates were incubated for 3 hours and then treated with gentamicin to kill the extracellular bacteria.
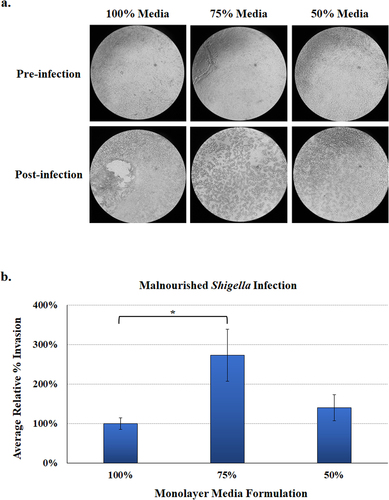
Table 1. Primers used in this study.
Data availability statement
The authors confirm that the data supporting the findings of this study are available within the article.
