Figures & data
Table 1. Characteristics of 112 children participating in a prospective cohort study of enteric infections in Villa El Salvador, Lima, Peru, 2016–2019.
Figure 1. Changes in children’s exposures to breastmilk over the first 16 months of life, Lima, Peru, 2016–2019. Children’s feeding patterns were derived from 52,816 daily survey visits.
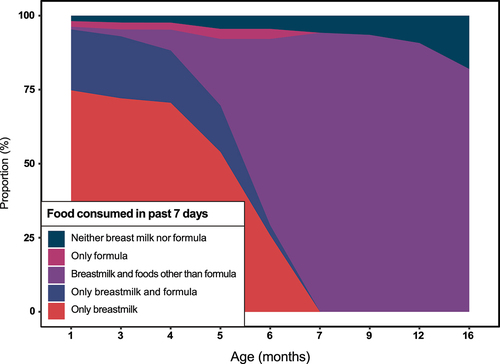
Table 2. Early-life feeding patterns of 112 children participating in a prospective cohort study of enteric infections in Villa El Salvador, Lima, Peru, 2016–2019.
Figure 2. Gut colonization with ESBL-producing Enterobacterales over the first 16 months of life, Lima, Peru, 2016–2019. Changes in the mean point prevalence and incidence of a) ESBL-producing Escherichia coli and b) ESBL-producing Klebsiella spp., Enterobacter spp., or Citrobacter spp. Are depicted across specific age categories. Gray bands represent 95% confidence intervals around the mean.
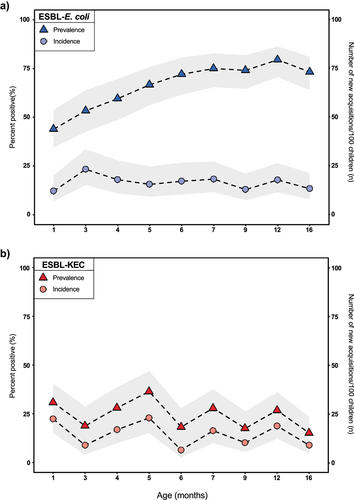
Table 3. Associations between breastfeeding over the first 16 months of life and children’s incident gut colonization with extended-spectrum β-lactamase-producing enterobacterales, Lima, Peru, 2017–2019.
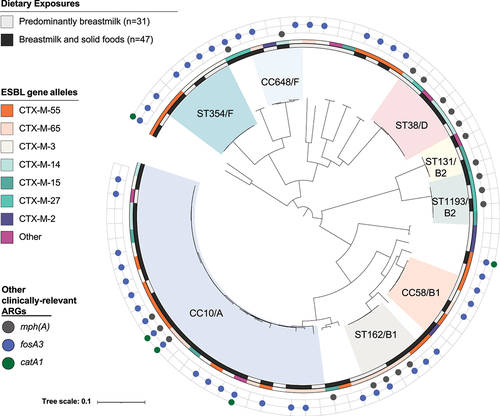
Figure 3. Core genome phylogenetic tree depicting the evolutionary relationships of 78 ESBL-producing E. coli from 47 children in Lima across two time points (predominantly breastfeeding versus breastfeeding while also eating complementary foods). We observed a high diversity of ESBL-ec and few differences in the occurrence of specific clonal complexes (CCs), sequence types (STs), phylogroups, ESBL alleles, or other antibiotic resistance genes (ARGs) across time points, suggesting the same ESBL-ec sequence types that were circulating among the broader community, in animals, and in the environment were also circulating within the household environment. Phylogroup designations A, B1, B2, or F are listed after highlighted CCs and STs.