Figures & data
Table 1. Primers for PCR and qRT-PCR.
Table 2. Upregulated genes in prtP-deficient mutant.
Table 3. Genes downregulated in prtP-deficient mutant.
Table 4. Genes upregulated in msp-deficient mutant.
Table 5. Genes downregulated in msp-deficient mutant.
Figure 1. Expression of msp and prtP determined using qRT-PCR. (a) Expression of msp. (b) Expression of prtP (dentilisin); 35405: T. denticola ATCC 35405 (wild type), K1: dentilisin mutant, DMSP3: msp mutant. The expression of each gene is presented as fold-change over that in the wild type strain. Data are presented as means ± SD (n = 6), and statistically significant differences are indicated using asterisks (one-way ANOVA with Dunnett’s multiple comparison test; *p < 0.05 compared to the wild type).
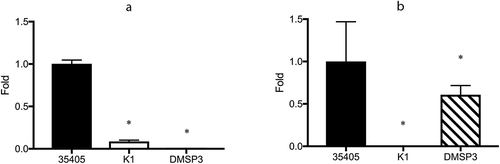
Figure 2. SDS-PAGE and immunoblot analysis of sonicates of T. denticola ATCC 35405, DMSP3, and K1. SDS-PAGE analysis (a) and immunoblot analysis (b).
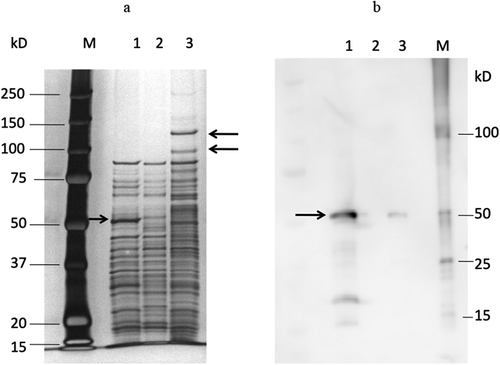
Figure 3. Dentilisin activity of wild type (35405) and msp-deficient mutant (DMSP3). Data are presented as means ± SD, and statistically significant differences are indicated by asterisks (student t-test; *p < 0.05 compared to the wild type). 35405: T. denticola ATCC 35405, DMSP3: T. denticola DMSP3, cell: cells of T. denticola, sup: culture supernatant of T. denticola, log: log phase, stationary: stationary phase.
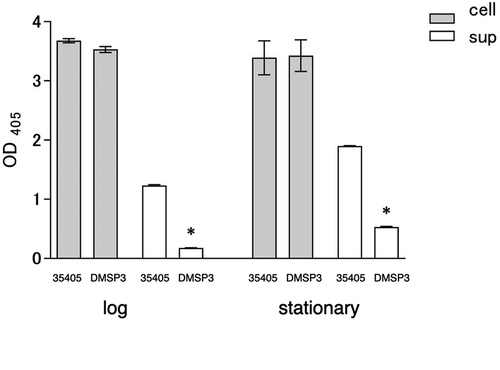
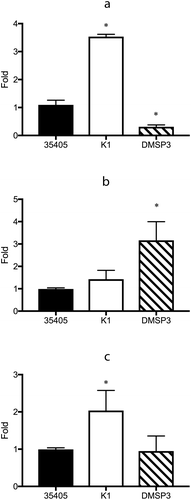
Figure 4. Gene expression of the potential regulator proteins in wild type (35405), msp-deficient mutant (DMSP3), and prtP-deficient mutant (K1). Gene expression levels were evaluated using qRT-PCR with a Taqman probe. The expression of TDE_0127 (a), TDE_0344 (b), and TDE_0814 (c) are presented as fold-change with respect to that of the wild type strain. Data are presented as means ± SD (n = 6), and statistically significant differences are indicated by asterisks (one-way ANOVA with Dunnett’s multiple comparison test; *p < 0.05 compared to wild type).