Figures & data
Table 1. Samples of two typical light-flavor Daqus of Chinese spirits.
Table 2. Bacterial diversity and amylase production in Niulanshan Daqu.
Table 3. Bacterial diversity and amylase production in Hongxing Daqu.
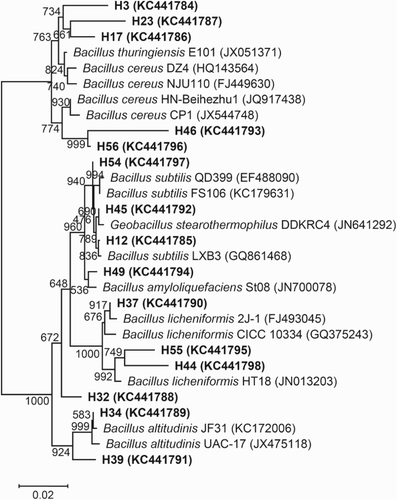
Figure 1. Phylogenetic relationships among bacterial 16S rDNA sequences in the Daqu of Niulanshan liquor and with previously reported sequences. The number on each branch indicates the percentage of 1000 replicates that are included in the branch. Sequences determined in this study are shown in bold, and GenBank accession numbers are shown in parentheses for all of the related sequences. The scale bar of 0.02 represents a 2% nucleotide substitution rate according to the Jukes–Cantor evolutionary distance (Surhio et al. Citation2014).
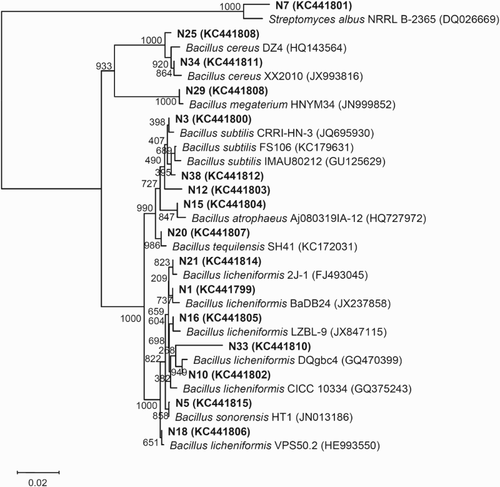
Figure 2. Phylogenetic relationships among bacterial 16S rDNA sequences in the Daqu of Hongxing liquor and with previously reported sequences. The number on each branch indicates the number out of 1000 replicates that are included in the branch. Sequences determined in this study are shown in bold, and GenBank accession numbers are shown in parentheses for all of the related sequences. The scale bar of 0.02 represents a 2% nucleotide substitution rate according to the Jukes–Cantor evolutionary distance.