Figures & data
Figure 1. Mechanisms of conjugative mobilization in Staphylococci. The conjugative plasmid encodes all genes required for formation of the mating pore, as well as the coupling protein, DNA relaxase and an oriT. Mobilizable plasmids can exploit the conjugative-plasmid mating pore by either: (A) encoding a mimic sequence of the conjugative-plasmid oriT; (B) encoding a distinct relaxase (Mob) compatible with the conjugative plasmids coupling protein and its own cognate oriT or; (C) by carrying a replicative relaxase (Rep) compatible with the conjugative-plasmid coupling protein.
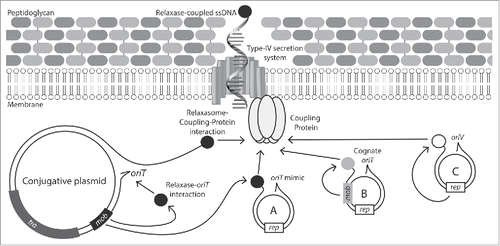
Figure 2. pWBG4, a third family of staphylococcal conjugative plasmids. The internal circle represents the gene map of pWBG4, showing the positions and predicted products of the putative pWBG4 conjugation cluster detA-detV (white arrows with black outlines) and other open-reading frames. The outer circles represent ungapped circular BLASTN alignments of pWBG4-family plasmids, created using BRIG software.Citation63
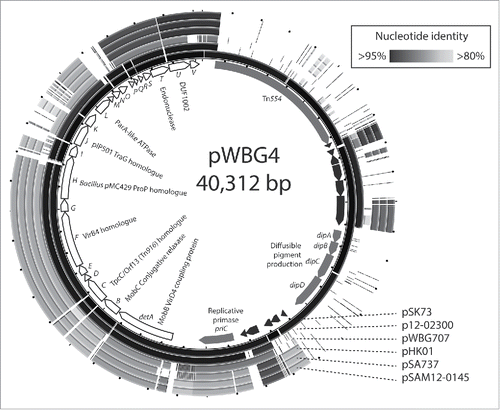
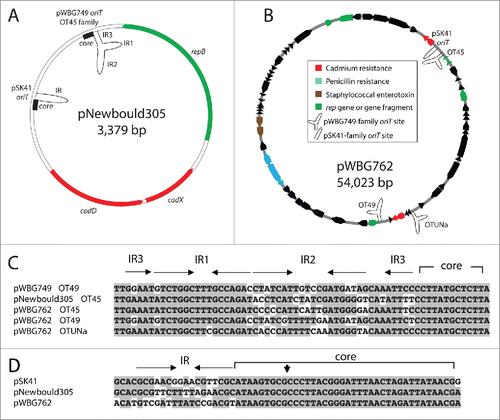
Figure 3. The diversity of oriT mimics on large and small resistance plasmids in staphylococci. (A) The plasmid map of the rolling-circle plasmid pNewbould305, illustrates the presence of 3 potential mobilization mechanisms. The repB gene of pNewbould305 shares 56% amino-acid identity over 96% of its length with the pBS42/pUB110 RepB protein, which enables mobilization by ICEBs1-family elements. Downstream of the repB gene is an oriT mimic sequence of the pWBG749-family, subfamily OT45 and a pSK41-like oriT mimic sequence. (B) The atypically large staphylococcal plasmid pWBG762, carries 4 oriT mimics. Three are of the pWBG749 family and one is of the pSK41 family. (C) Alignment of the pWBG749-family oriT mimic sequences carried by pNewbould305 and pWBG762 below the pWBG749 oriT region, illustrating IR2 sequence divergence. Conserved nucleotides are shaded. The AR1-AR3 repeats of the full oriT required for mobilization by pWBG749Citation8 have been truncated in this figure for clarity (D) Alignment of the pSK41-like oriT mimic sequences from pNewbould305 and pWBG762, below the pSK41 oriT region, showing divergence of the IR sequence; the Nes relaxase nick site is denoted by a vertical arrowhead.