Figures & data
Figure 1 Reconstitution of STAT1 in Ifnar1−/− MEFs partially rescues IFNγ responses. (A) Protein lysates from wild type, Ifnar1−/−, Ifnar1−/−MSCV and Ifnar1−/−HA-STAT1 129/C3H MEFs were separated by SDS-PAGE and probed with antibodies specific for STAT1 and actin as a loading control. (B-G) wild type, Ifnar1−/−, Ifnar1−/−MSCV and Ifnar1−/−HA-STAT1 129/C3H MEFs were treated with 100 U/ml IFNγ for 1, 3 and 6 hrs and mRNA expression was assessed by Affymetrix microarray (n = 3). (B-D) Venn diagram of the number of genes that are up-regulated (↑ green) or down-regulated (↓ red) relative to untreated (p < 0.05, >1.2-fold) following (B) 1 hr, (C) 3 hrs or (D) 6 hrs of IFNγ treatment and whether they are specific or common to each cell type. (E-G) Left panel. Heat maps of probes with attenuated IFNγ-dependent gene induction in Ifnar1−/− MEFs relative to wild type MEFs (p > 0.05, <1.2-fold change relative to untreated or >1.2-fold attenuation in change compared to wild type). Right panel. Heat maps of probes with attenuated (as defined above) IFNγ-dependent gene induction in Ifnar1−/− MEFs and Ifnar1−/−MSCV MEFs relative to wild type MEFs and rescued (p < 0.05 and >1.2-fold change relative to untreated and <1.2-fold attenuation in change compared to wild type) in Ifnar1−/−HA-STAT1 MEFs. The 10 attenuated (left panel) and rescued (right panel) probes with the greatest fold change in wild type MEFs following (E) 1 hr, (F) 3 hrs and (G) 6 hrs of IFNγ treatment are displayed. Heatmaps were generated using Java Treeview. Total number of probes with induced (↑) and repressed (↓) expression in wild type MEFs that had attenuated (left panel) or rescued (right panel) expression are listed below heatmaps.
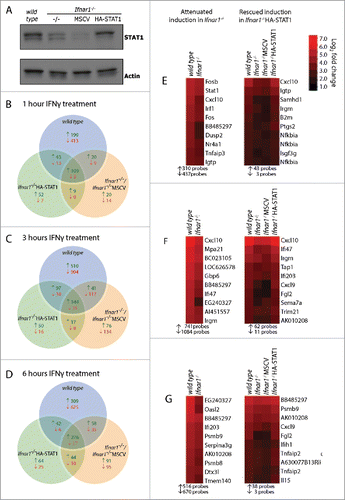
Figure 2 Reconstitution of STAT1α in Ifnar1−/− MEFs preferentially rescues immune and inflammatory IFNγ responses. (A-B) Probes from listed as attenuated (A-B) or rescued (B) were analyzed using MetaCore's Genego. The 10 most significantly affected pathways are displayed. (A) Pathway enrichment analysis of attenuated genes. EMT = epithelial-to-mesenchymal transition. (B) Comparative pathway enrichment analysis of attenuated and rescued genes. (C) Heatmaps displaying fold change in expression following 1, 3 or 6 hrs IFNγ treatment of genes involved in antigen presentation via MHC class I for wild type, Ifnar1−/−, Ifnar1−/−MSCV and Ifnar1−/−HA-STAT1 MEFs. Heatmaps were generated using Java Treeview.
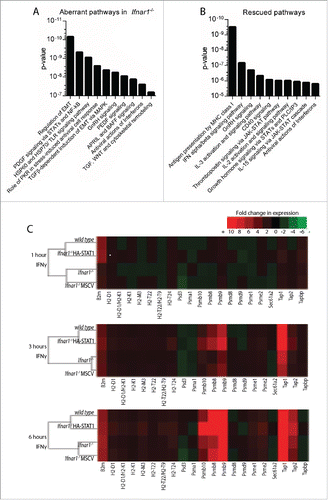
Figure 3. STAT1 reconstitution rescues IFN-induced protein expression of MHC class I antigen presentation proteins in Ifnar1−/− MEFs. (A-C) wild type, Ifnar1, Ifnar1−/−MSCV and Ifnar1−/−HA-STAT1 MEFs were cultured in media alone or treated with 100 U/ml IFNγ for (A) 6 hrs and (B-C) 24 hrs. Protein lysates were separated by SDS-PAGE and probed with antibodies specific for (A) TAP1 (B) TAP2 or (C) PSMB9. Western membranes were re-probed with antibody specific for (A-B) Actin or (C) HSP70 to confirm equivalent loading of proteins. (D-E) wild type, Ifnar1, Ifnar1−/−MSCV and Ifnar1−/−HA-STAT1 MEFs were cultured in media alone or were treated with 5 or 10 U/ml IFNγ for 16 hrs. Cells were analyzed by flow cytometry for expression of the MHC class I molecules (D) H2-Kb and (E) H2-Db. Data represents the IFNγ-induced change in MFI relative to untreated for 11–13 independent experiments and error bars are SEM. Significant changes in MFI relative to untreated (fold increase in MFI) were determined by t-test #p < 0.05, ##p < 0.01, ###p < 0.001. Significant differences in fold increase in MFI relative to wild type were determined by t-test *p < 0.05, **p < 0.01, ***p < 0.001.
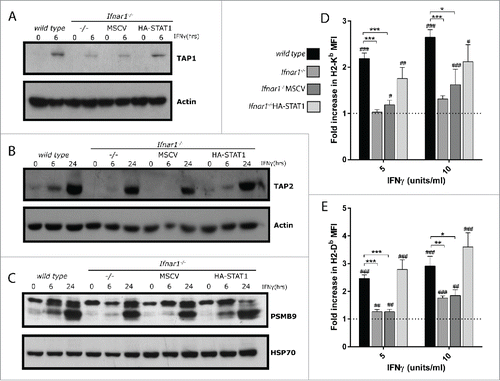
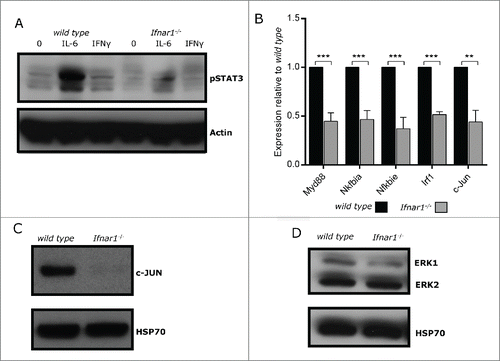
Figure 4. Aberrant expression and activation of signaling intermediaries in unstimulated Ifnar1−/− MEFs. (A) wild type and Ifnar1−/− MEFs were cultured in media alone or treated with 100 U/ml IFNγ or 100 ng/ml IL-6 and 100ng/ml IL-6R for 30mins. Protein lysates were separated by SDS-PAGE and probed with an antibody specific for pSTAT3. Western membrane was re-probed with an antibodies specific for Actin to confirm equivalent loading of proteins. (B) RNA was extracted from untreated wild type and Ifnar1−/− 129/C3H MEFs and mRNA expression of c-Jun, Irf1, Myd88, Nfkbiα and Nfkbie was determined by qRT-PCR. Expression of each gene was normalized to L32 and untreated wild type MEFs. Data represents the mean of 4–6 independent experiments and error bars are SEM. Significance determined by t-test *p < 0.05, **p < 0.01, ***p < 0.001. (C-D) Protein lysates from untreated wild type and Ifnar1−/−MEFs were separated by SDS-PAGE and probed with antibodies specific for (C) c-JUN, and (D) ERK1/2. Western membranes were re-probed with antibodies specific for HSP70 to confirm equivalent loading of proteins.