Figures & data
Figure 1. Mutational landscape of significantly mutated genes in triple-negative breast cancer (TNBC) patients.
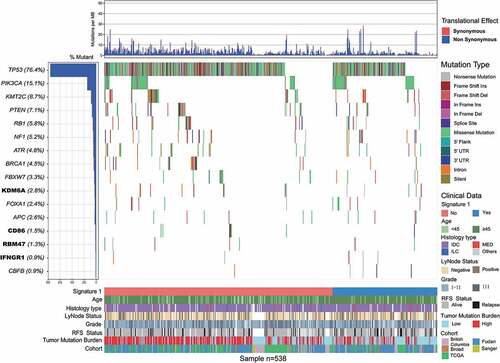
Figure 2. Mutational signatures extracted from the aggregated TNBC dataset.
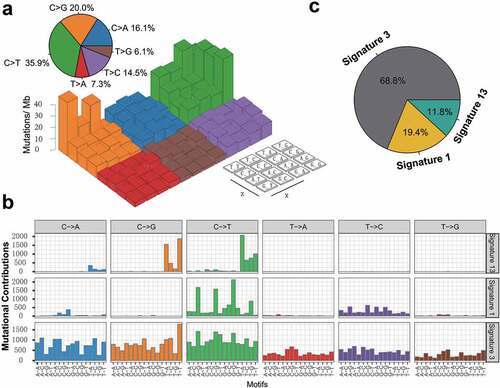
Figure 3. Association of mutational signature 1 with prognosis in TNBC cohort.
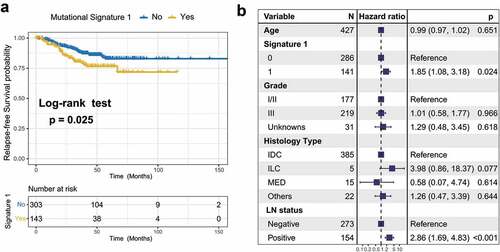
Figure 4. Significantly enriched pathways and immune infiltration alteration with signature 1 activities.
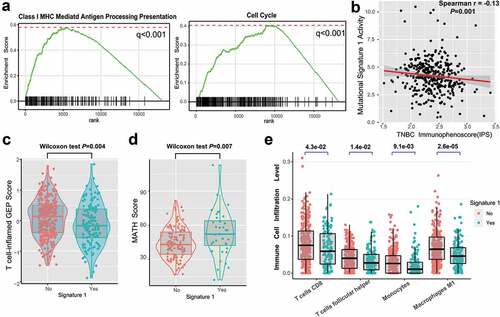
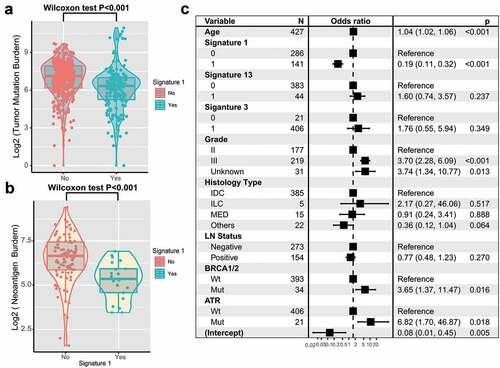
Figure 5. Association of signature 1 with tumor mutation burden in pooled TNBC cohort.