Figures & data
Figure 1. Chemiluminescence counts reflecting reactive oxygen species (ROS) production from PMNs challenged with Pseudomonas aeruginosa vaccine strains (PAO1, PAO3, PAO5, PAO6, PAO9, and PAO11). The chemiluminescence, given in counts per second (CPS), was measured using luminol in a microplate fluorescence reader (Wallac 1420 Victor2, Perkin Elmer) over 60 minutes of phagocytosis. Each panel shows the peak CPS ± SEM of experiments run in duplicate from 3 different donors. *Lowest concentration of S-IgY to give a significant increase in respiratory burst compared to the control (p < 0.05)
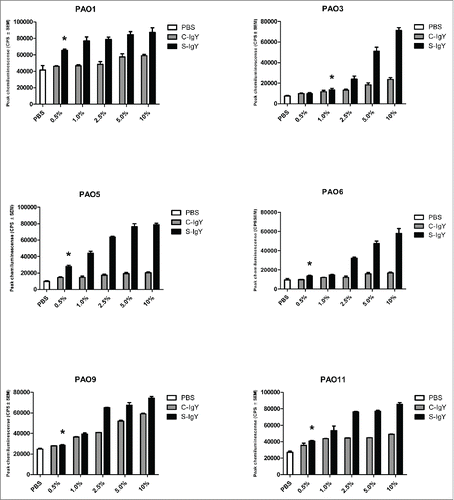
Figure 2. Chemiluminescence counts reflecting reactive oxygen species (ROS) production from PMNs challenged with non-mucoid Pseudomonas aeruginosa clinical strains (516/08, 5762C/07, 39014/06, 40542A/07, and 40542B/07) isolated from 4 non-chronically infected CF patients and a flagella mutant (PAO1172 TTN 183). The chemiluminescence, given in counts per second (CPS), was measured using luminol in a microplate fluorescence reader (Wallac 1420 Victor2, Perkin Elmer) over 60 minutes of phagocytosis. Each panel shows the peak CPS ± SEM of experiments run in duplicate from 3 different donors. *Lowest concentration of S-IgY to give a significant increase in respiratory burst compared to the control (p < 0.05)
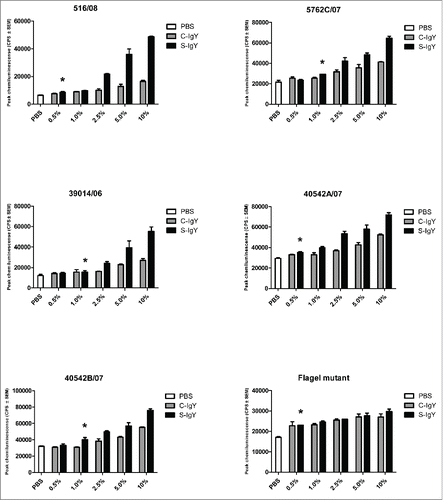
Figure 3. Chemiluminescence counts reflecting reactive oxygen species (ROS) production from monocytes challenged with the PAO1 vaccine strain. (A) depicts the respiratory burst level from monocytes phagocytizing 107 CFU/ml PAO1, which corresponds to a cell-to-bacteria ratio of 1:1. (B) shows the burst level from monocytes phagocytizing 108 CFU/ml PAO1, which corresponds to a cell-to-bacteria ratio of 1:10. The chemiluminescence, given in counts per second (CPS), was measured using luminol in a microplate fluorescence reader (Wallac 1420 Victor2, Perkin Elmer) over 60 minutes of phagocytosis. Each panel shows the peak CPS ± SEM of experiments run in duplicate from 3 different donors
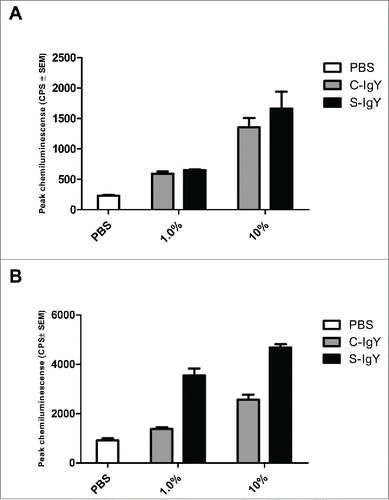
Figure 4. PMN-mediated bacterial killing depicted as the percentage of viable bacteria after 60 minutes of phagocytosis. PMNs were challenged with Pseudomonas aeruginosa vaccine strains (PAO1, PAO3, PAO5, PAO6, PAO9, and PAO11) and a flagella mutant (PAO1172 TTN 183), and the number of viable bacteria was determined by colony counting the next day. Each panel shows the % survival ± SEM of experiments run in duplicates. Significance was tested by one way ANOVA test with * representing comparison with PMN and non-IgY (-IgY) with p value < 0.05
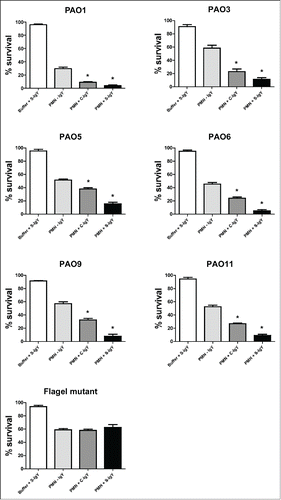
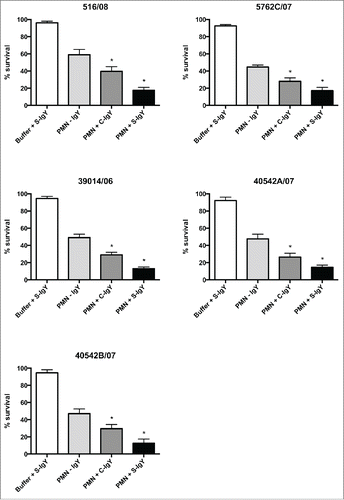
Figure 5. PMN-mediated bacterial killing depicted as the percentage of viable bacteria after 60 minutes of phagocytosis. PMNs were challenged with Pseudomonas aeruginosa clinical strains, and the number of viable bacteria was determined by colony counting the next day. Each panel shows the % survival ± SEM of experiments run in duplicates. Significance was tested by one way ANOVA test with * representing comparison with PMN and non-IgY (-IgY) with p value < 0.05