Figures & data
Figure 1. Analysis of purified IgG by SDS-PAGE and immunoblot analysis of mAb secreted by hybridomas. (A) Analysis of purified antibody by SDS-PAGE. Lanes 1, 2: purified mAb M-H11; Lane M: molecular mass markers (B) Immunoblot analysis of mAb secreted by hybridomas. Lane 1: purified fusion protein reacted with mAb M-H11; Lane M: molecular mass markers.
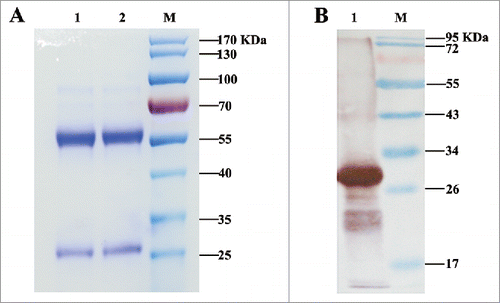
Figure 2. Analysis and characterization of Mpep. (A) Detection of the binding activity of the phage clones by ELISA. 1–32: phage clones from the third round of biopanning; B: BSA control. (B) The sequences of 12 peptides displayed by 15 phage clones. Multiple sequence alignment was performed using Clustal Omega. The symbol * means the site has the same amino acid, the symbol. means the amino acids belong to one group. Glutarnine (Q) and Serine (S) are both of uncharged polar side chains group. The blue box represents the same amino acid. (C) Affinity reaction of Mpep and M-H11 (Epep as control). Data were expressed as the mean ± standard deviation (S.D). The experiment was repeated 3 times. (D) Competitive inhibition of Mpep against MraY-AB and M-H11.
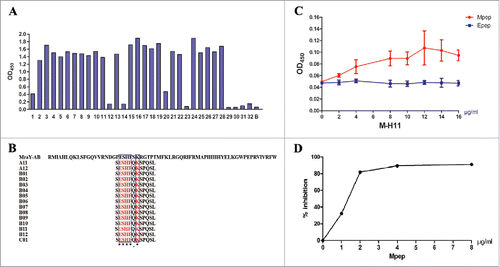
Figure 3. M-H11 has the ability to inhibit the E. coli BL21 (DE3) plysS growth in vitro and in vivo. (A) Line chart showing M-H11 inhibition of E. coli BL21 (DE3) plysS growth in vitro. (B) Histogram showing the inhibition rate of M-H11 on E. coli BL21 (DE3) plysS growth in vitro. (C) Histogram showing the inhibition ability of M-H11 on E. coli BL21 (DE3) plysS growth in vivo. Six-week-old female BALB/c mice were randomly divided into 2 groups (5 in each group) in vivo experiment.
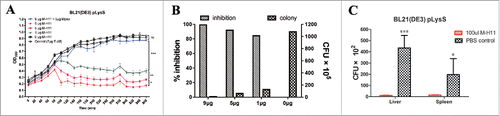
Table 1. The colony count of organs in mice infected with E. coli BL21 (DE3) plysS alone.
Table 2. Primers used in this study.
