Figures & data
Figure 1. (a) Illustration of SARS-CoV RBD subunit S1. (b) Sequence alignment between SARS-CoV RBD219-N1 and SARS-CoV-2 spike protein. The RBM region is circled in green. An example of a neutralizing conformational epitope consisting S343–367, 373–390 and 411–428 (reported by Bian et al.) is circled in blue, indicating neutralizing epitopes do not have to be within RBM.Citation7
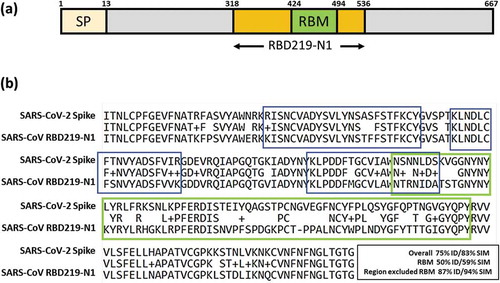
Table 1. Neutralizing monoclonal antibodies reported in He et al, 2005Citation11 were categorized based on its ability to inhibit RBM binding to the ACE2.
Table 2. Binding study of anti-SARS-CoV RBD neutralizing mAb against the SARS-CoV-2 RBD
