Figures & data
Table 1. Primers for qRT-PCR.
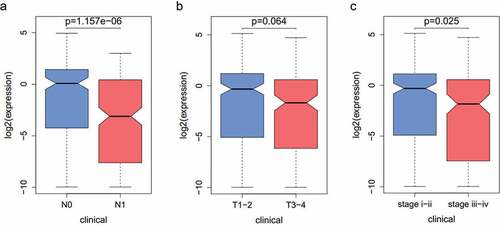
Figure 1. CXCL5 expression in TC. (a) CXCL5 expression in TC tissues. the blue and pink box plots represent the normal samples and the tumor samples, respectively; (b) the effects of CXCL5 on the prognosis of TC. (c) mRNA expressions of CXCL5 in normal cell line (Nthy-ori3–1) and TC cell lines (KTC-1, TPC-1, BHP10–3, K1, BCPAP); (d) protein expressions of CXCL5 in different cell lines (*p < .05).
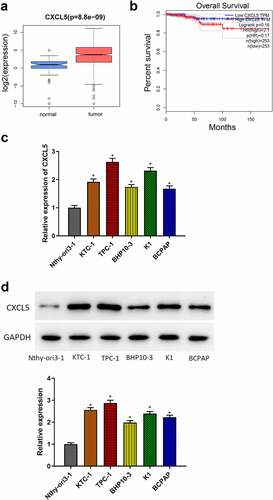
Figure 2. CXCL5 promotes malignant behaviors of TC cells. (a) CXCL5 transfection efficiency of each group; (b) proliferative capacity of cells in each group; (c) cell proliferation in each group; (d) cell migratory ability in each group; (e) cell invasive ability in each group (*p < .05).
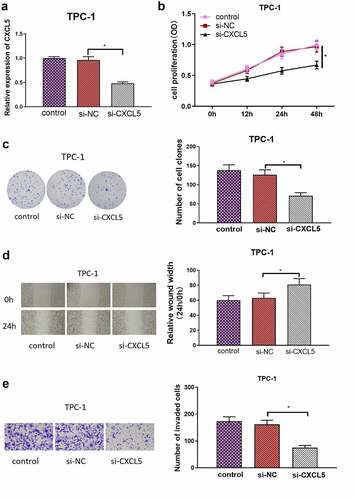
Figure 3. CXCL5 is a direct target of microRNA-873-5p. (a) Venn diagram of predicted microRnas and down-regulated differential microRnas; (b) Pearson correlation analysis of microRNA-873-5p and CXCL5; (c) microRNA-873-5p expression in TC tissues. The blue and pink box represent the normal samples and the tumor samples, respectively; (d) MicroRNA-873-5p expression in TC cells; (e) Targetscan predicts upstream regulatory genes of CXCL5; (f) Verification of the binding of microRNA-873-5p with CXCL5; (g) the expression level of CXCL5 mRNA in cells with microRNA-873-5p overexpression; (h) the expression level of CXCL5 protein in cells with microRNA-873-5p overexpression (*p < .05).
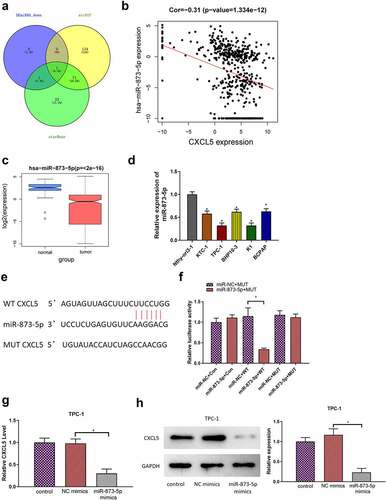
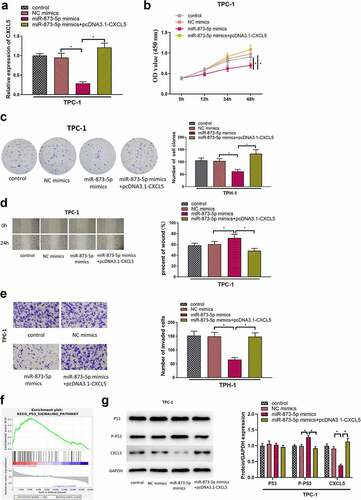
Figure 4. MicroRNA-873-5p mediates p53 pathway by targeting CXCL5 to inhibit cell proliferation, migration, invasion in TC. (a) The transfection efficiency of each group; (b) Cell viability in each group; (c) Cell proliferation in each group; (d) Cell migratory ability in each group; (e) Cell invasive ability in each group; (f) GSEA of CXCL5; (g) the expression levels of CXCL5 protein and P53-related pathway proteins (*p < .05).
Data availability statement
All data generated or analyzed during this study are included in this published article.