Figures & data
Table 1. The benchmark dataset S taken from Virus-mPlocCitation21.
Figure 1. (a) The impact of each residue on the subsequent residues. . The impact of each residue on the subsequent residues. . The impact of each residue on the subsequent residues.
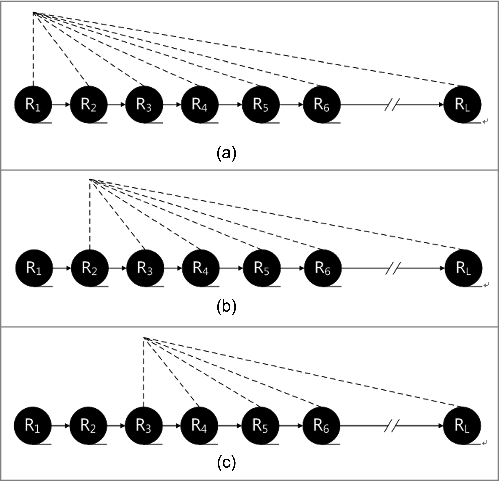
Table 2. Sorting signals of database.
Table 3. Application of two methods to original database and new database.
Table 4. Six physicochemical properties.
Table 5. Application of PseAAC, R-Dipeptide, I-PseAAC and PseAAC2 to the new database.
Table 6. Overall accuracy of R-Dipeptide, I-PseAAC, and PseAAC2.
Table 7. Evaluation functions for PseAAC, R-Dipeptide, I-PseAAC, and PseAAC2.
