Figures & data
Table 1. Feeding deterrence index (FDI) of R. dominica exposed to C. procera extracts-treated food at sub-lethal concentrations, t = 3 days.
Table 2. (a): Observed and predicted FDI against extracts concentrations using Probit, Logit, Log-log, Clog-log, Cauchy, Fractional, and Optimal models, t = 3 days. (b): Observed and predicted FDI against extracts concentrations using Probit, Logit, Log-log, Clog-log, Fractional, Cauchy and Optimal models, t = 3 days.
Table 3. ε and for all used extracts with Probit, Logit, Log-log, Clog-log, Cauchy and fractional models.
Table 4. The parameters a, b for Probit, Logit, Log-log, Clog-log, Cauchy and fractional models.
Table 5. Concentrations corresponding to 25%, 50% and 75% FDI using Probit, Logit, Log-log, Clog-log, and Cauchy models.
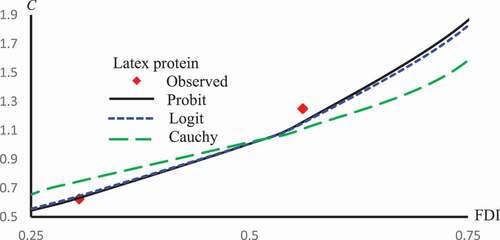
Figure 2. Observed and predicted FDI for various concentrations of Latex protein using Probit, Logit and Cauchy models.
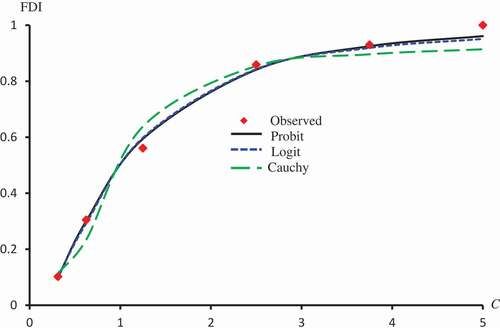
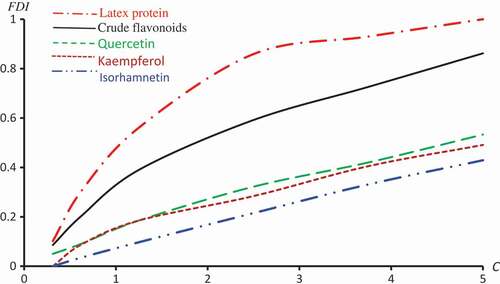
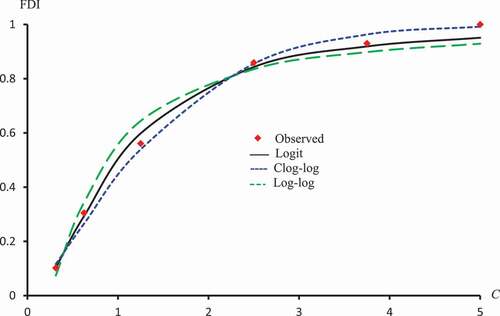
Figure 3. Observed and predicted FDI for various concentrations of Crude flavonoids using Probit, Clog-log Cauchy and Fractional models.
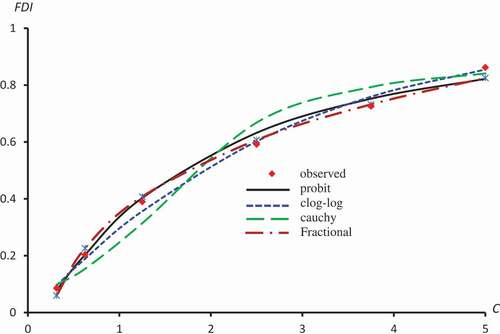
Figure 4. Observed and predicted FDI for various concentrations of Quercetin-3-O-rutinoside using Probit, Clog-log, Cauchy and Fractional models.
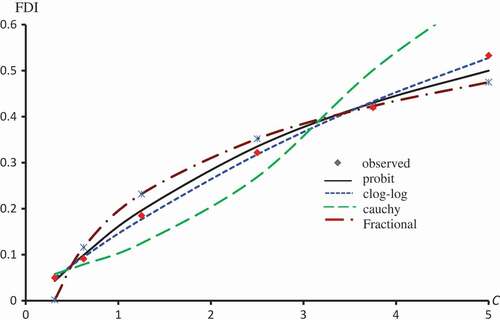
Figure 5. Observed and predicted FDI for various concentrations of Kaempferol-3-O-rutinoside using Probit, Clog-log and Cauchy models.
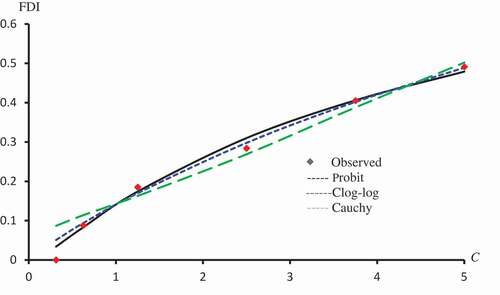
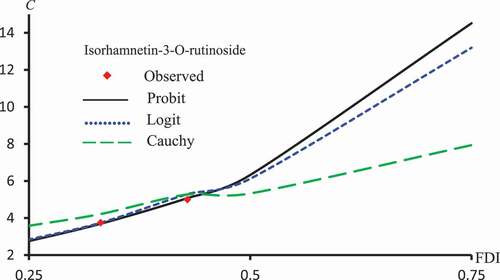
Figure 6. Observed and predicted FDI for various concentrations of Isorhamnetin-3-O-rutinoside using Probit, Logit and Cauchy models.
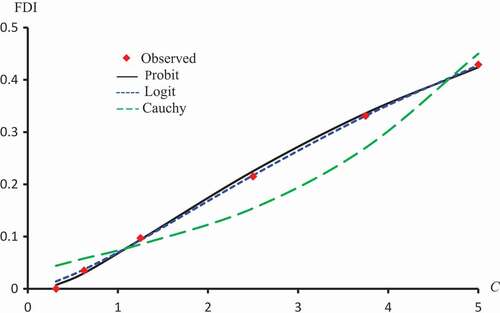
Figure 7. Observed FDI, predicted FDI for minimum and predicted FDI with
,
for various concentrations of Latex protein using Probit models.
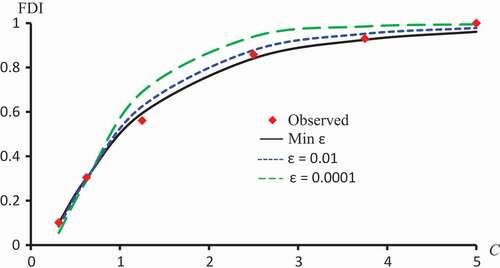
Figure 8. Observed FDI, predicted FDI for minimum and predicted FDI with
and
for various concentrations of Kaempferol-3-O-rutinoside using Logit models.
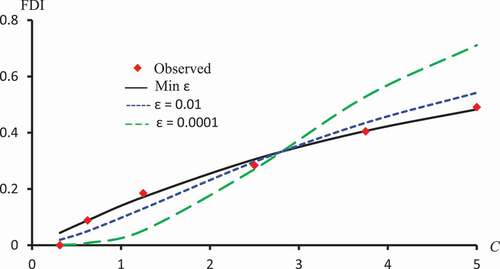
Figure 9. Observed FDI, predicted FDI for minimum and predicted FDI with
=0.0001 and
for various concentrations of Isorhamnetin-3-O-rutinoside using Clog-log models.
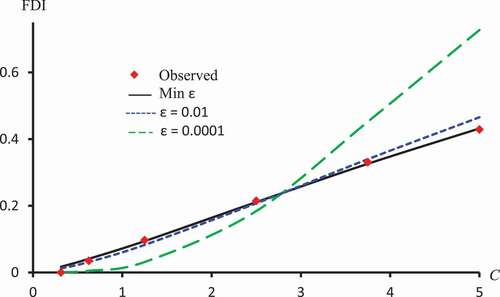
Figure 10. against ε for Latex protein, Kaempferol-3-O-rutinoside and Isorhamnetin-3-O-rutinoside using Probit models.
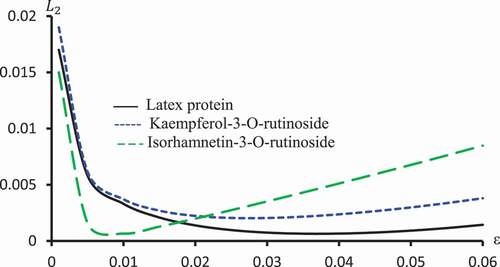
Figure 11. Observed and predicted latex protein concentrations corresponding to FDI using Probit, Logit and Cauchy models.