Figures & data
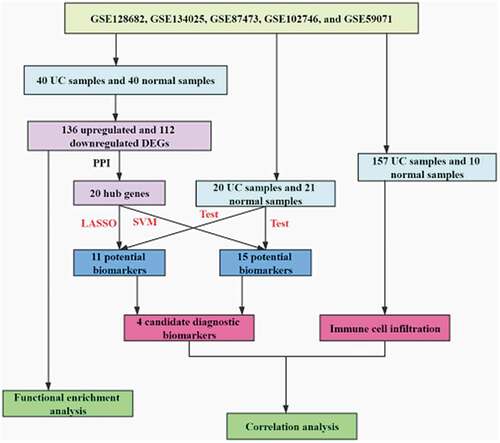
Figure 1. Data collection, processing, and DEG identification
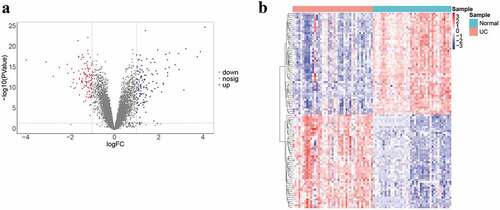
Figure 2. GO and KEGG analysis
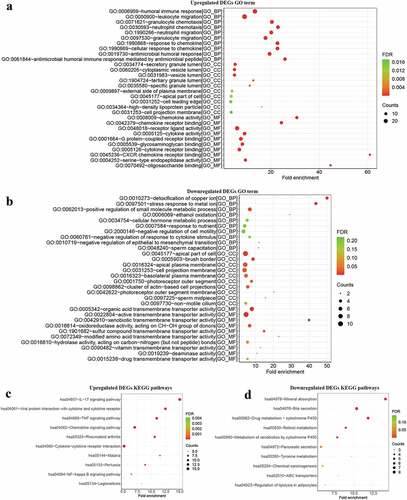
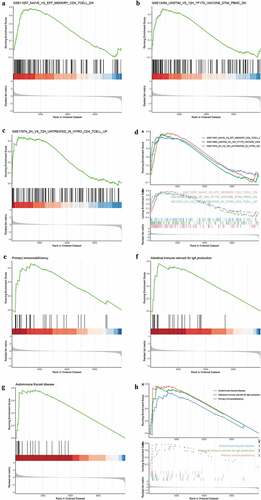
Figure 5. Identification and validation of candidate UC biomarkers
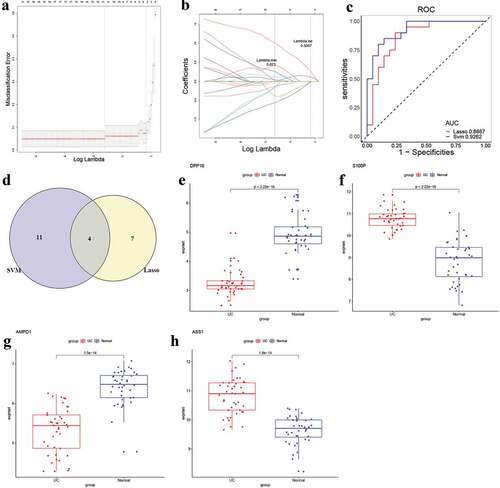
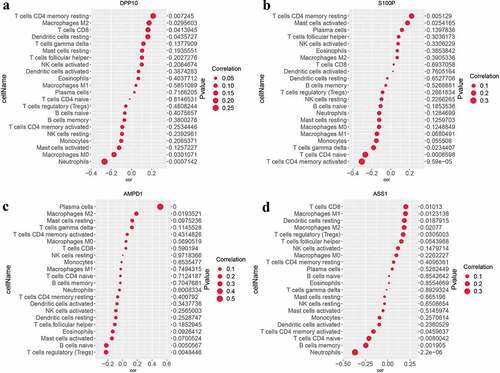
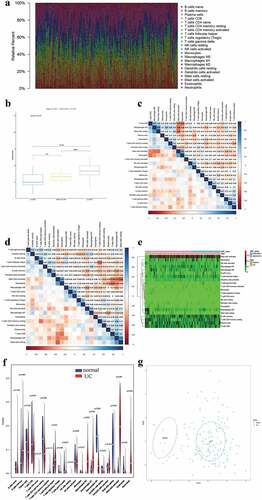
Figure 6. Assessment of immune infiltration
Supplemental Material
Download ()Data availability statement
All of the datasets analyzed were acquired from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/gds).