Figures & data
Table 1. The correlation between hsa_circ_0001287 and clinicopathological features of NSCLC patients
Figure 1. The expression of circ_0001287 in NSCLC
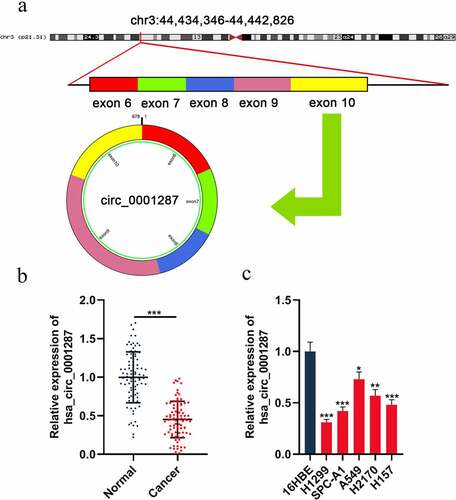
Figure 2. Circ_0001287 inhibited the multiplication, migration, invasion, and radioresistance of NSCLCs
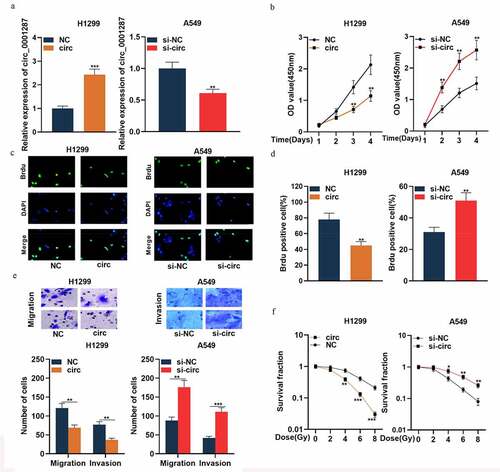
Figure 3. Circ_0001287 adsorbed miR-21
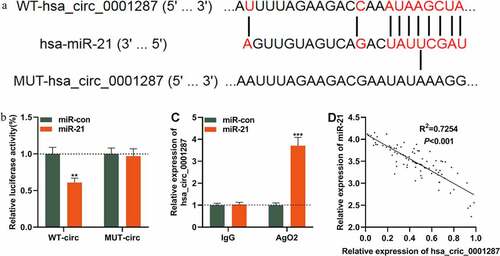
Figure 4. MiR-21 played an oncogenic role in NSCLCs
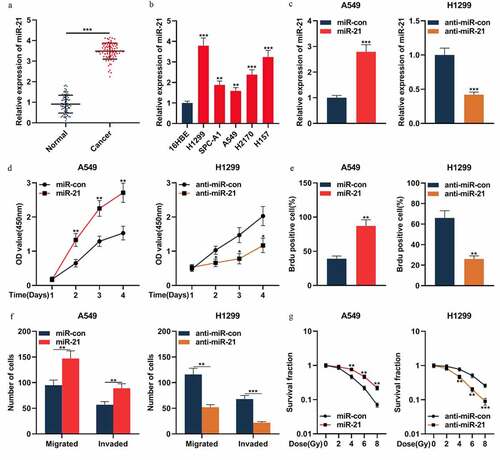
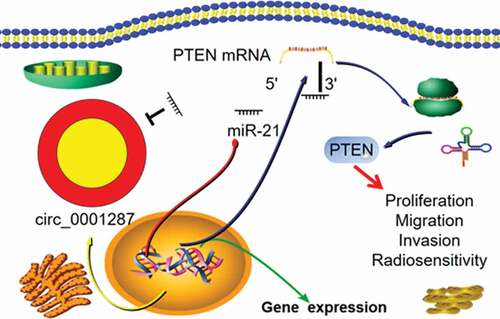
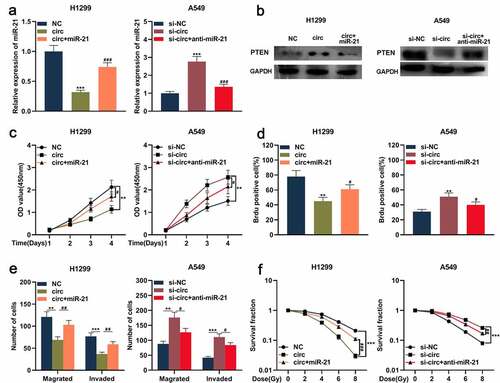
Figure 5. Circ_0001287/miR-21/PTEN axis was involved in regulating the malignant phenotype of NSCLCs
Supplemental Material
Download ()Data Availability Statement
The data used to support the findings of this study are available from the corresponding author upon request.