Figures & data
Figure 1. Cisatracurium inhibits the viability of OVCAR-3 cells and enhances the expression of p53 and lincRNA-p21 in a dose-dependent manner. (a) OVCAR-3 cells were treated with different doses (10, 20 and 40 μM) of cisatracurium for 24, 48 and 72 h, and cell viability was analyzed by CCK-8 assay. (b) Expression of p53 was determined in the OVCAR-3 cells using WB. (c) Expression of p53 was determined in the OVCAR-3 cells using real-time PCR. **p < 0.01 and ***p < 0.001 vs. Control. n = 3
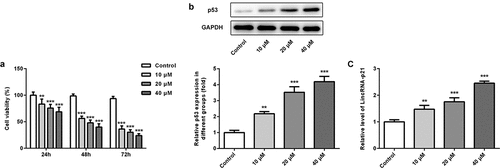
Figure 2. Cisatracurium inhibits the proliferation of OVCAR-3 cells by activating lincRNA-p21. (a) Overexpressed efficiency of p53 expression was detected by real-time PCR. ***p < 0.001 vs. Control. (b) Knockdown efficiency of p53 expression was detected by real-time PCR. ***p < 0.001 vs. Control. (c) LincRNA-p21 expression in OVCAR-3 cells transfected with Ov-p53 or shRNA-p53 was detected by real-time PCR. ***p < 0.001 vs. Control; ###p < 0.001 vs. Ov-NC; ∆∆∆p < 0.001 vs. shRNA-NC. (d) Knockdown efficiency of lincRNA-p21 expression was detected by real-time PCR. (e) Cell proliferation was determined by CCK-8 assay. (f) Cell proliferation was determined by Edu staining. ***p < 0.001 vs. Control; ###p < 0.001 vs. 20 μM+sh-NC. n = 3
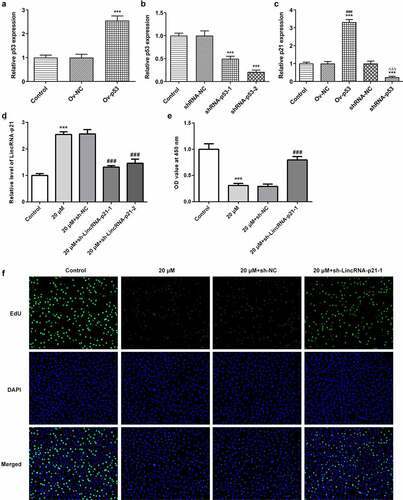
Figure 3. Cisatracurium inhibits migration and invasion of OVCAR-3 cells by activating lincRNA-p21. (a and b) Cell migration was determined by wound-healing assay. (c and d) Cell invasion was determined by Transwell assay. (e) The expression of MMP2, MMP9, TIMP1 and TIMP2 were measured by WB. ***p < 0.001 vs. Control; #p < 0.05, ##p < 0.01 and ###p < 0.001 vs. 20 μM+sh-NC. n = 3
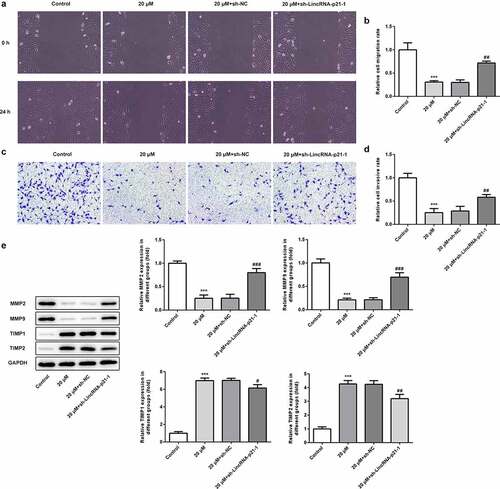
Figure 4. Cisatracurium promotes apoptosis of OVCAR-3 cells by activating lincRNA-p21. (a) Cell apoptosis were determined by TUNEL staining. (b) The expression levels of apoptosis-related proteins (Bax, Bcl-2, caspase3 and Cleaved caspase3) were determined by WB. ***p < 0.001 vs. Control; ###p < 0.001 vs. 20 μM+sh-NC. n = 3
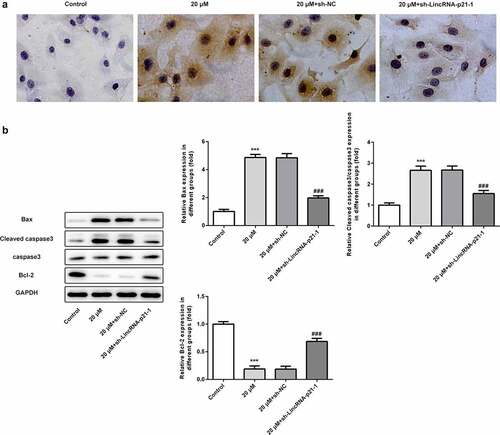
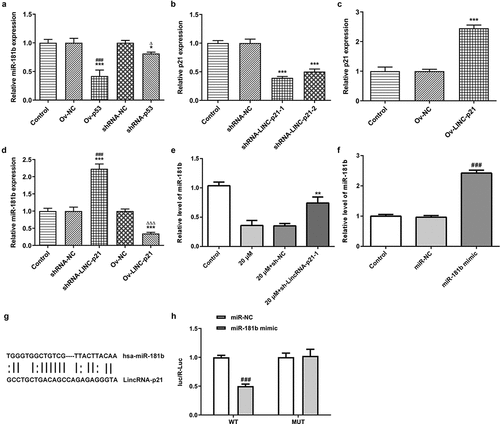
Figure 5. Cisatracurium inhibits the expression of miR-181b by activating lincRNA-p21. (a) miR-181b expression in OVCAR-3 cells transfected with Ov-p53 or shRNA-p53 was detected by real-time PCR. *p < 0.05 and ***p < 0.001 vs. Control; ∆p < 0.05 vs. shRNA-NC. (b) Knockdown efficiency of lincRNA-p21 expression was detected by real-time PCR. ***p < 0.001 vs. Control. (c) Overexpressed efficiency of lincRNA-p21 expression was detected by real-time PCR. ***p < 0.001 vs. Control. (d) miR-181b expression in OVCAR-3 cells transfected with shRNA-LINC-P21 or Ov-LINC-P21 was detected by real-time PCR. ***p < 0.001 vs. Control; ###p < 0.001 vs. shRNA-NC; ∆∆∆p < 0.001 vs. Ov-NC. (e) Effect of lincRNA-p21 on miR-181b expression was determined by real-time PCR. **p < 0.001 vs. 20 μM+sh-NC. (f) Overexpression efficiency of miR-181b expression was detected by real-time PCR. ###p < 0.001 vs. miR-NC. (g) Predicted binding sites for miR-181b in lincRNA-p21. (h) The relationship between lincRNA-p21 and miR-181b was determined by luciferase reporter assay in OVCAR-3 cells. n = 3