Figures & data
Figure 1. STK39 was up-regulated in HCC and predicts poor prognosis
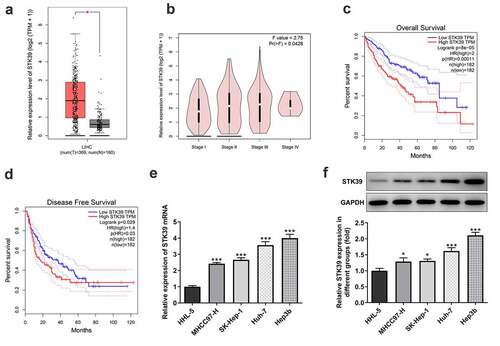
Figure 2. STK39 knockdown inhibited proliferation and induces apoptosis of Hep3b cells
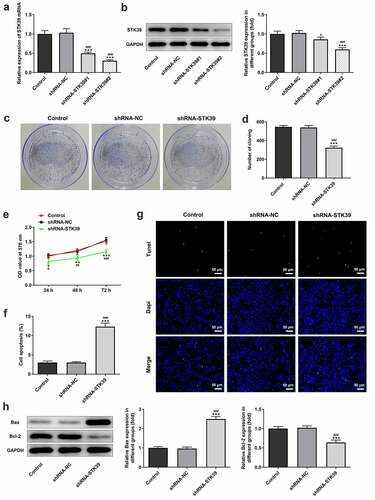
Figure 3. STK39 knockdown suppressed migration, invasion, EMT and TGF-β1/Smad2/3 signaling of Hep3b cells
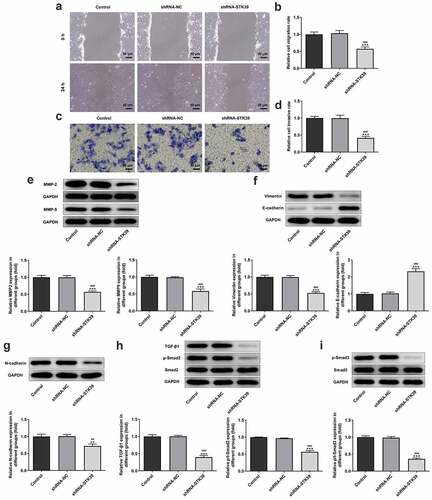
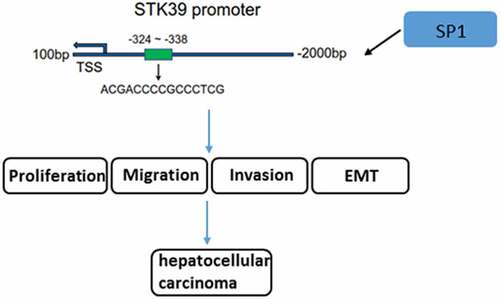
Figure 4. The relationship between SP1 and STK39
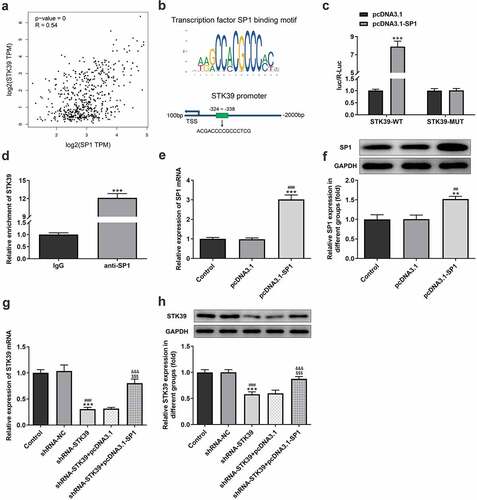
Figure 5. SP1 overexpression blocked the effect of STK39 knockdown on Hep3b cells proliferation and apoptosis
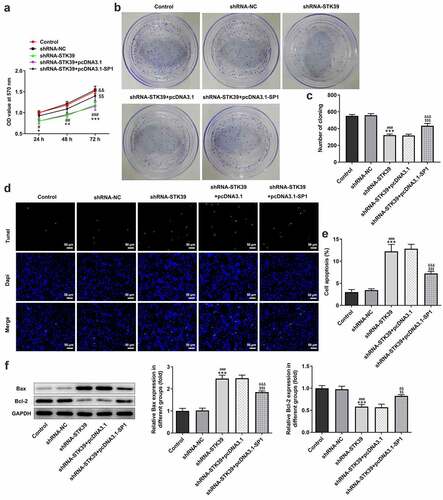
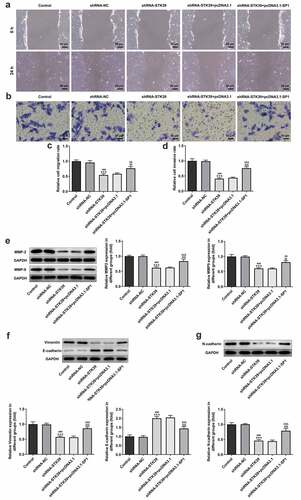
Figure 6. SP1 overexpression alleviated the effect of STK39 knockdown on Hep3b cells migration, invasion and EMT
Availability of data and materials
The datasets generated and/or analyzed during the current study are available on reasonable request.