Figures & data
Table 1. Correlation between circRNA_102231 expression and clinicopathological features in GC patients (n = 120)
Figure 1. CircRNA_102231 is upregulated in GC. (a). qRT-PCR analysis of circRNA_102231 expression in paired GC and normal tissues. (b). The survival curve of GC patients based on median circRNA_102231 level in GC tissues. (c, d). qRT-PCR analysis of plasma circRNA_102231 expression, followed by ROC curve analyzing its diagnostic value. ***p< 0.001. ANT = adjacent normal tissue
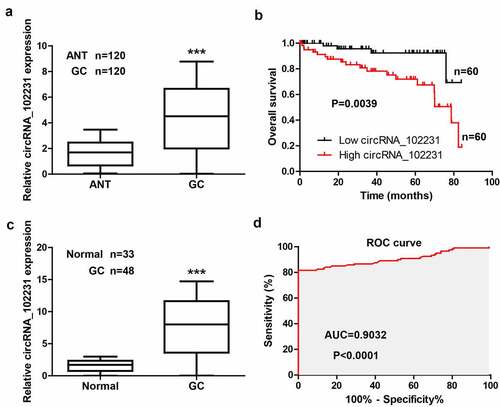
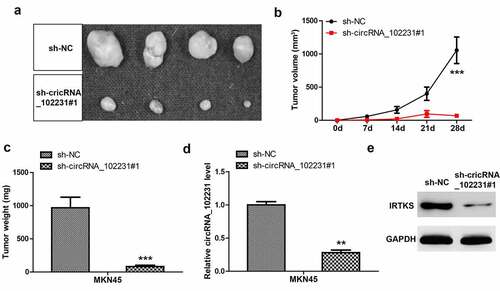
Figure 2. Knockdown of circRNA_102231 inhibits GC progression. (a, b). qRT-PCR analyzing the knockdown efficiency of these siRNAs. (c–j). Colony formation, EdU, CCK-8 and Transwell assays in GC cells after circRNA_102231 knockdown. *p< 0.05, **p< 0.01
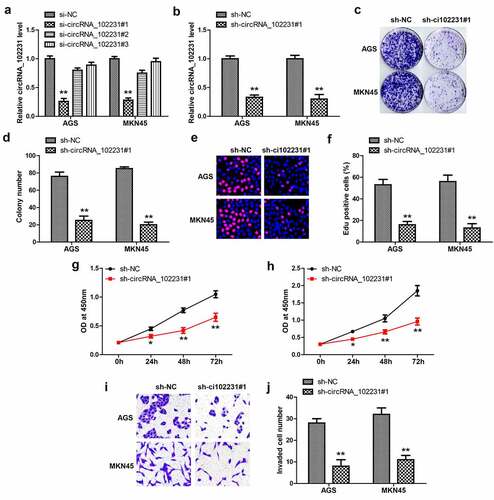
Figure 3. CircRNA_102231 interacts with IRTKS. (a, b). qRT-PCR analysis of the location of circRNA_102231. C. The top ten proteins pulled down by circRNA_102231. (d–f). qRT-PCR verifying the specificity and effectiveness of circRNA_102231 probe, followed by western blot assay. G, H. RIP assay testing the enrichment of circRNA_102231 by anti-IRTKS. **p< 0.01
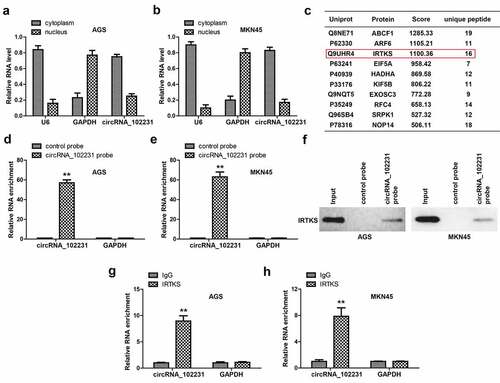
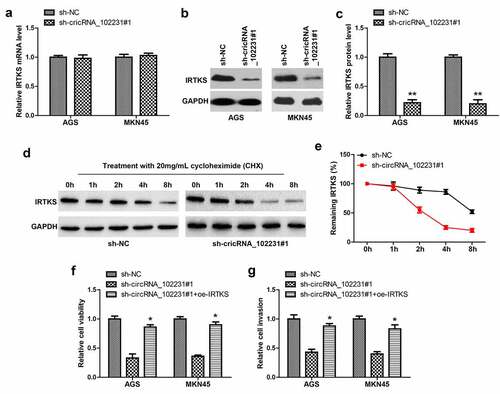
Figure 4. CircRNA_102231 increases IRTKS protein stability. (a). qRT-PCR analysis of IRTKS mRNA level in circRNA_102231-silenced GC cell lines. (b, c). Western blot testing IRTKS protein in circRNA_102231-silenced GC cell lines. (d, e). Western blot testing the effect of circRNA_102231 on IRTKS protein stability. (f, g). Cell viability and invasion assay in circRNA_102231-silenced GC cell lines transfected with IRTKS-expressing vector. *p< 0.05, **p< 0.01