Figures & data
Table 1. Demographic distribution of GDI1 in a pooled dataset of colon cancer
Table 2. Uni- and multivariate analysis for GDI1 and survival in CRC datasets
Table 3. Demographic distribution of colorectal cancers from ZJU set
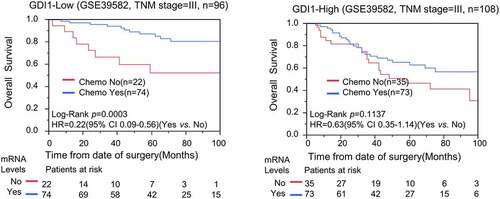
Figure 1. Distribution of the GDI1 mRNA expression and aggressiveness of CRC
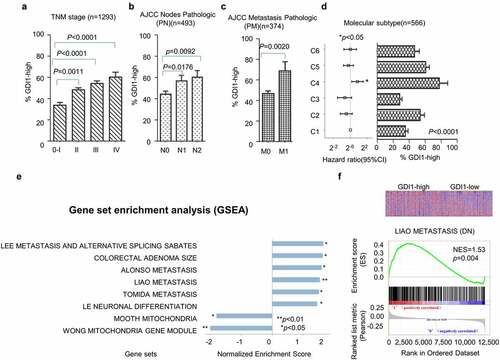
Figure 2. Survival analysis for GDI1 mRNA expression and outcome of CRC
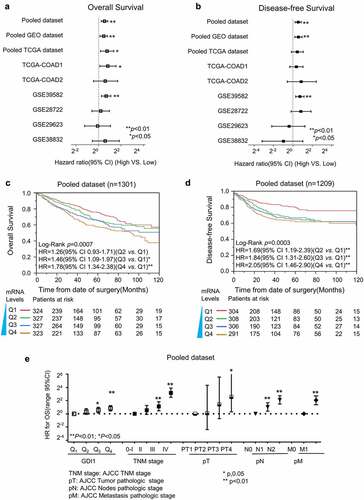
Figure 3. Immunohistochemistry analysis for the protein expression of GDI1 in the cytoplasm and membrane
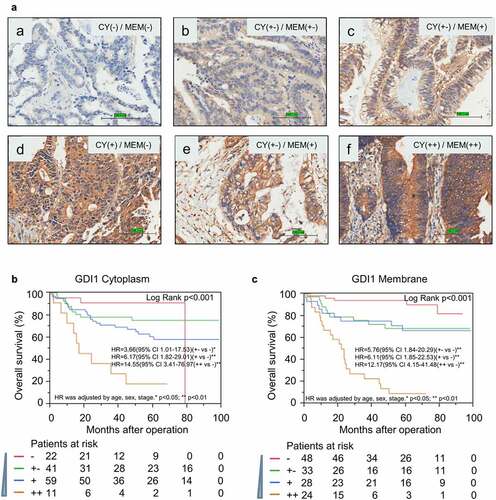
Figure 4. Stratification analysis for GDI1 expression and chemotheresistance in CRC patients