Figures & data
Figure 1. Expression of hsa_circ_0075541 and hsa_circ_0075542 in prostate cancer tissues. (a and b): hsa_circ_0075542 and hsa_circ_0075541 expression levels in prostate cancer and adjacent tissues, respectively; n = 30. (c): Sequence and (d): gel electrophoresis image of the quantitative reverse transcription polymerase chain reaction amplification products of hsa_circ_0075541 and hsa_circ_0075542
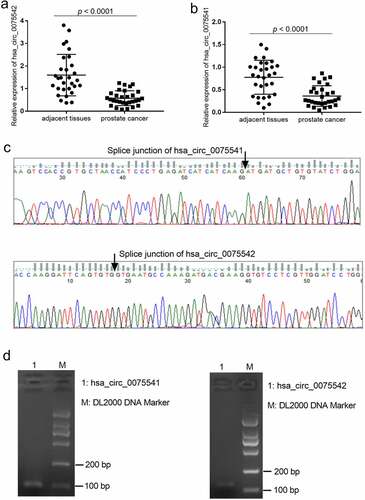
Figure 2. hsa_circ_0075542 plays a tumor-suppressive role in LNCaP and PC3 cells. LNCaP and PC3 cells were transfected with overexpression of hsa_circ_0075542 vector (ov-circ_0075542) or empty vector pLC5-ciR (empty vector). (A): hsa_circ_0075542 level in the above-mentioned transfected cells was determined using quantitative reverse transcription PCR. B-E: The effect of this transfection on cell proliferation (B) and apoptosis (C) was assessed using cell counting kit-8 and flow cytometry assays. The migration (D) and invasion (E) capabilities were evaluated using transwell assays. *P < 0.05, the ov-circ_0075542 group vs. the empty vector group
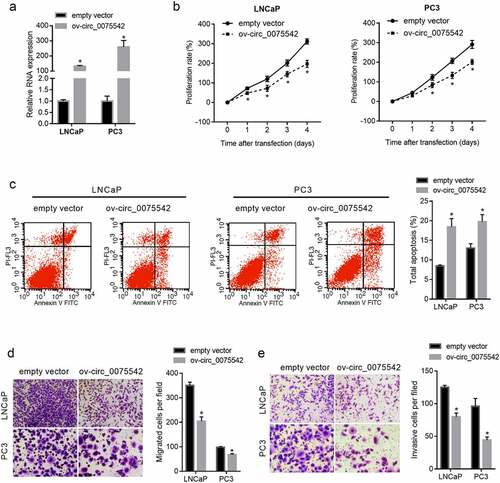
Figure 3. hsa_circ_0075542 targets the miR-1197/HOXC11 axis. (A): miR-1197 binding sites on hsa_circ_0075542 and 3′-noncoding region of HOXC11 mRNA. (B): Effect of miR-1197 mimic or miR-NC on the luciferase activity of recombinant luciferase reporter. To construct a luciferase reporter, the wild-type linear sequence of hsa_circ_0075542 was cloned into the miRNA target expression vector GP-mirGLO, named wild-type mirGLO-circ_0075542. miR-1197 binding site on wild-type mirGLO-circ_0075542 was mutated and the mutant plasmid was named mutant mirGLO-circ_0075542. miR-NC or miR-1197 was co-transfected with each recombinant luciferase reporter plasmid, and the luciferase activity was measured. (C): Relative expression of hsa_circ_0075542 in pulled-down RNA using biotinylated miR-NC or biotinylated miR-1197 as a probe. The ‘input’ group is the isolated RNA from the partial lysate before the cell pull-down assay. (D): HOXC11 protein level in cells transfected with overexpression of hsa_circ_0075542 vector (ov-circ_0075542) or empty vector pLC5-ciR (empty vector). (E): HOXC11 protein abundance in LNCaP and PC3 cells transfected with miR-NC, miR-1197 mimic, miR-1197 mimic plus empty vector, or miR-1197 mimic plus ov-circ_0075542. *P < 0.05
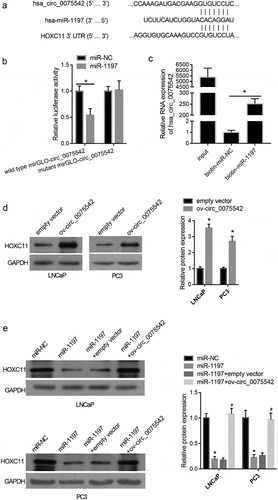
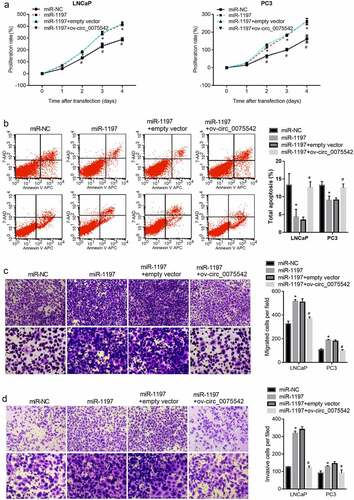
Figure 4. Overexpression of hsa_circ_0075542 alleviated the tumor-promoting effect of miR-1197 overexpression. After being transfected with miR-NC, miR-1197 mimic, miR-1197 mimic plus empty vector, or miR-1197 mimic plus ov-circ_0075542, the effect of this transfection on cell proliferation (a) and apoptosis (b) was assessed using cell counting kit-8 and flow cytometry assays. The capabilities of migration (c) and invasion (d) were evaluated using transwell assays. *P < 0.05, when miR-1197 group vs. the miR-NC group; #P < 0.05, when miR-1197 + ov-circ_0075542 group vs. the miR-1197 + empty vector group
Supplemental Material
Download ()Availability of data and materials
All data from this study are available in this published article.
