Figures & data
Table 1. Sequences used for qRT-PCR
Figure 1. SNHG18 expression in glioma and its biological function in regulating cell proliferation.
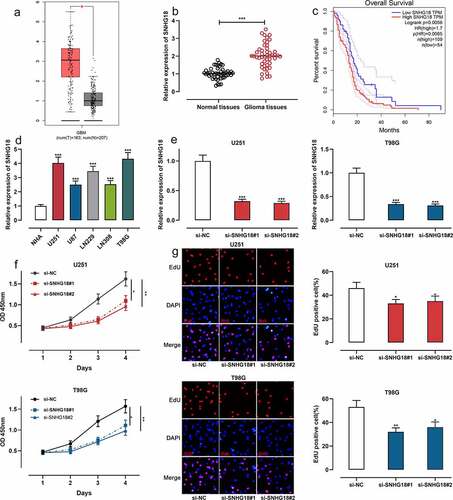
Table 2. Relationship between clinicopathological features and expression of SNHG18 in glioma
Figure 2. Knockdown of SNHG18 inhibits the migration and invasion of glioma.
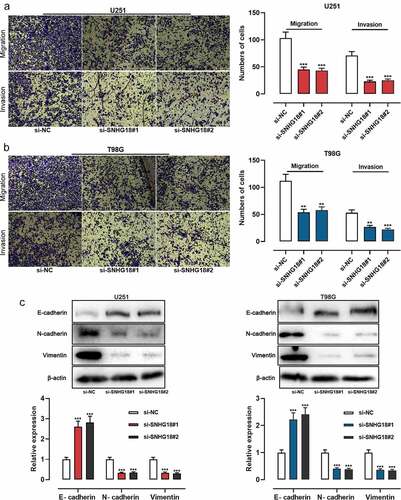
Figure 3. SNHG18 specifically regulates miR-338-5p expression.
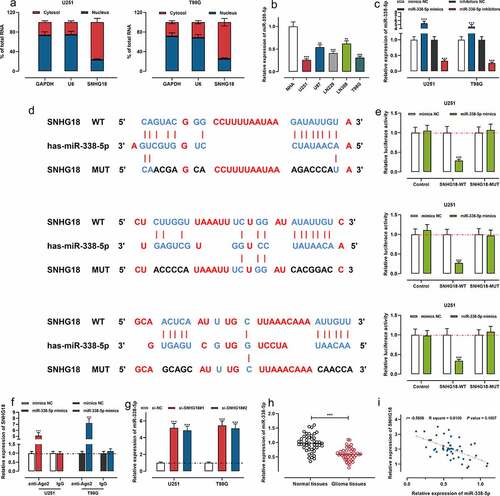
Figure 4. MiR-338-5p inhibitors can reverse the effects of SNHG18 knockdown on the proliferation, migration, and invasion of U251 and T98G cells.
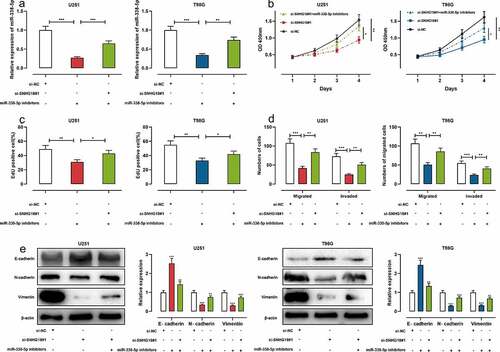
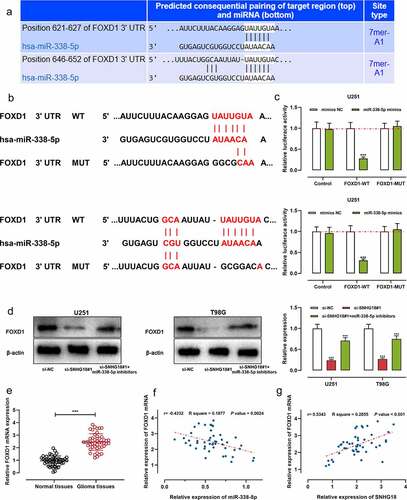
Figure 5. SNHG18 knockdown can inhibit FOXD1 expression.
Supplemental Material
Download Zip (17.9 MB)Data Availability Statement
The data used to support the findings of this study are available from the corresponding author upon request.
