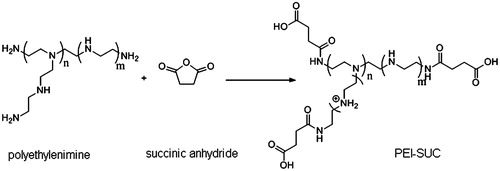
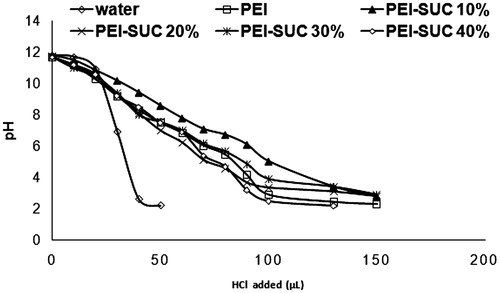
Figures & data
Table 1. Degree of primary amine conjugation in 25 kDa PEI estimated and size and zeta potential of the plasmid/DNA polyplexes.
Figure 1. Plasmid DNA condensation by 25 kDa PEI and its conjugated derivatives determined by the ethidium bromide exclusion assay.
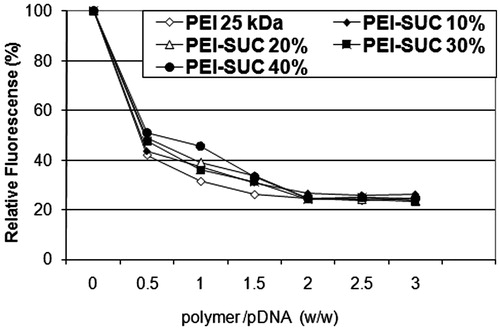
Figure 2. Gel retardation assay of unmodified PEI and its derivatives at various carrier/plasmid (C/P) ratios. Lanes 1, 6, and 11: unmodified 25 kDa PEI/pDNA at C/P ratios of 0.25, 4, and 8, respectively; Lanes 2, 7, and 12: PEI-SUC 10%/pDNA at C/P ratios of 0.25, 4, and 8, respectively; Lanes 3, 8, and 13, PEI-SUC 20%/pDNA at C/P ratios of 0.25, 4, and 8, respectively; Lanes 4, 9, and 14, PEI-SUC 30%/pDNA at C/P ratios of 0.25, 4 and 8, respectively; Lanes 5, 10, and 15, PEI-SUC 40%/pDNA at C/P ratios of 0.25, 4, and 8, respectively. Lane 16: DNA ladder.
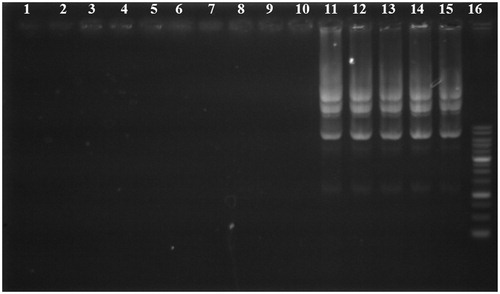
Figure 3. DNase I protection assay. Lanes 1–4: PEI-SUC 10% series treated with DNase I at C/P ratios of 0.25, 2, 4, and 8, respectively; Lanes 5–8: PEI-SUC 20% series treated with DNase I at C/P ratios of 0.25, 2, 4, and 8, respectively; Lanes 9–12: PEI-SUC 30% series treated with DNase I at C/P ratios of 0.25, 2, 4, and 8, respectively; Lanes 13–16: PEI-SUC 10% series treated with DNase I at C/P ratios of 0.25, 2, 4, and 8, respectively; Lane 17: DNA size marker.
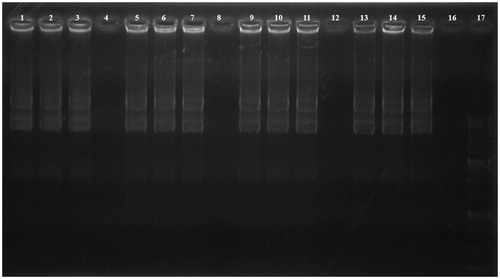
Figure 5. Hemolytic activity of unmodified PEI and its derivatives. 100% hemolysis was obtained using triton X-100 (final concentration of 0.1% w/v).
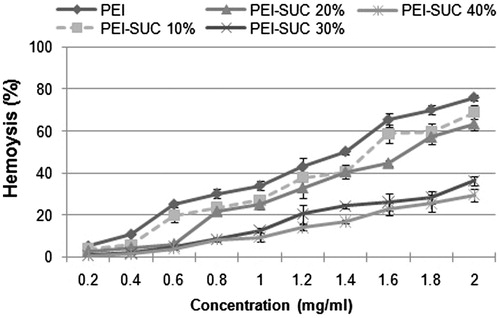
Figure 6. Cytotoxicity of unmodified 25 kDa PEI and its conjugates complexed with pUMVC3-hIL12 plasmid at C/P ratios of 0.25, 4, and 8 determined in triplicate in HepG2 cells. Metabolic activity was assayed using MTT method and expressed as the percentages of cell viability. *P < 0.05, conjugated PEI compared to unmodified parent polymer at the same C/P ratio.
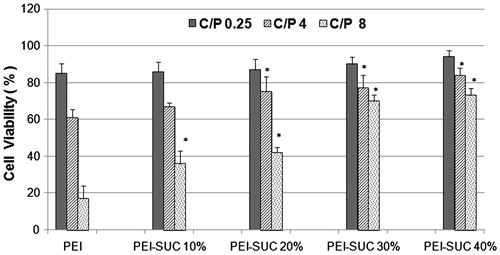
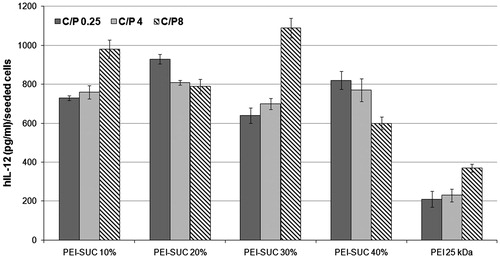
Figure 7. Transfection efficiency of unmodified and modified PEIs complexed with plasmid DNA at polymer:pDNA (weight:weight) ratios of 0.25, 4 and 8 determined in triplicate in HepG2 cell cultures. Cells were treated with polymer/pUMVC3-hIL12 polyplexes and ELISA was carried out to determine the level of hIL-12 production.