Figures & data
Figure 1. An illustrative structure of proposed T-nano-liposome. Liposomes can be surface functionalized to endow stealth through PEGylation and to promote receptor-mediated endocytosis by using targeting ligands such as pollen antigens, SCFV, aptamer, and MIP. PEGylation extends liposomal circulation half-life in vivo by reducing clearance, immune recognition, and the non-specific absorption of serum proteins. Polyethylene glycol (PEG) density determines its structure at the liposome surface.
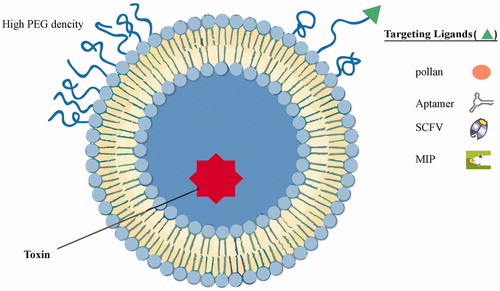
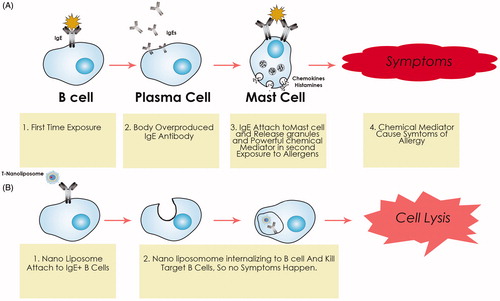
Figure 2. Schematic representation of the steps involved in the nano-liposomes-based target toxicity machine.
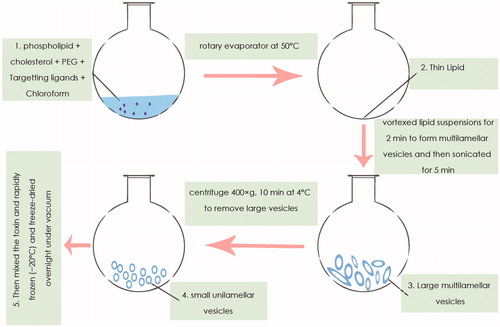
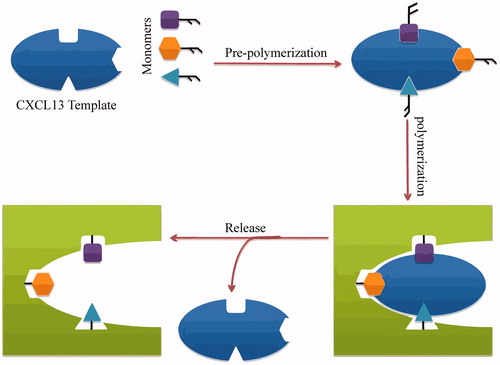
Figure 3. Scheme of molecular imprinting against CXCl 13. CXCL13 receptors are chosen as a target and an imprint molecule for this hypothesis. The first step is to consider the CXCL13 monomers and template (template is a constant, short peptide sequence illustrative of an available fragment of a larger protein, holes on the template is representative of the active sites in the CXCL13 template). Afterward, all elements of the MIPs are combined and allowed to be self-assembled to form the cross-linked polymer with the template (pre-polymerization). After polymerization, monomers, and the surrounding matrix are cleaved from the template molecules. The resulting targeted MIPs will be able selectively to bind to CXCL13.