Figures & data
Figure 1. (A) TEM image of CdNPs (right) and negatively stained CdNPs@BSA (left). (B) Schematic representation for synthesis of CdNPs@BSA. (C) Particle-size distribution histogram of CdNPs, mean diameter = 2 nm (P.D.I = 0.44) and (D) Schematic representation for synthesis of CdNPs@BSA. (E) Particle-size distribution histogram of free BSA, mean diameter = 7.5 nm (P.D.I = 0.38).
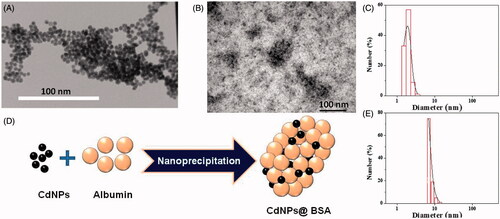
Figure 2. DLS plot of CdNPs@BSA, (A) First-order electric field correlation functions versus time at the indicated scattering angles. Inset plot: τ−1 as a function of q2. (B) First-order electric field correlation functions versus q2t at the indicated scattering angles. The inset plot shows random distribution and small values of the residuals. (C) First-order electric field correlation functions and their corresponding fits at 90°.
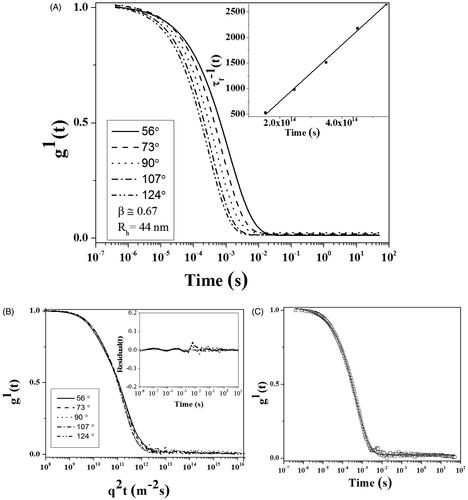
Figure 3. (A) UV–Vis spectra of CdNPs (▲▲▲▲), CdNPs@BSA (______) and BSA (- - -). (B) Emission fluorescence spectra of CdNPs@BSA (______), NAs (….), BSA (- - -). Far CD-UV (C) and near CD-UV (D) spectra of CdNPs@BSA (______), NAs (….) and BSA (- - -). Insert table: Secondary structural changes of BSA, NAs and CdNPs@BSA.
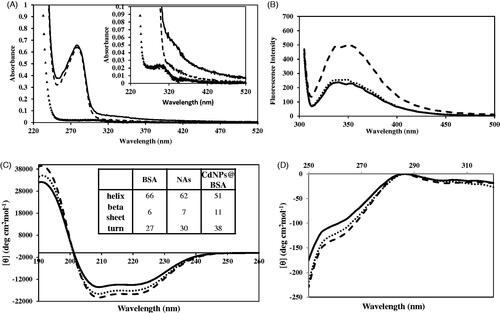
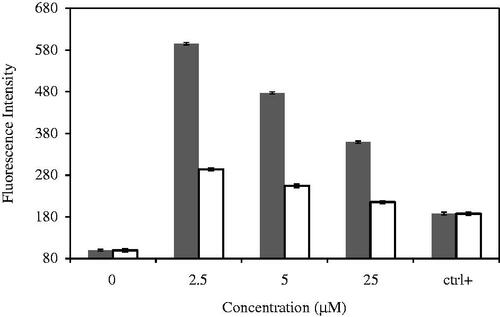
Figure 4. Cell viability of (A) MDA-MB 231, (B) WBC, (C) MCF-10A treated with CdNPs ▪ and CdNPs@BSA □. (D) Comparing cell viability of different kinds of cells treated with CdNPs and CdNPs@BSA.
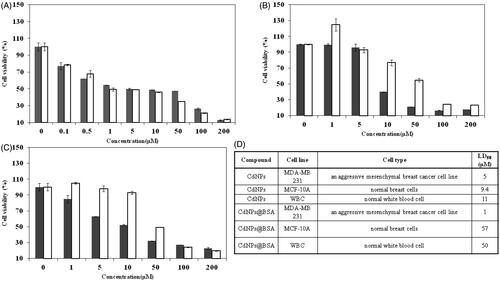
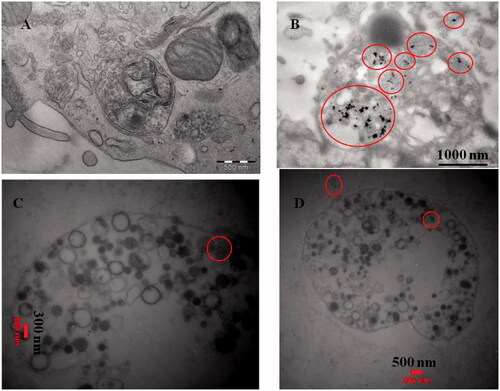
Figure 5. (UP) Inverted microscopic images of MDA-MB-231 Cells. (A) control, (B) and (C) treated with LD50 concentrations of CdNPs and CdNPs@BSA for 24 h, respectively. Middle) Fluorescent microscopy of cells stained with acridine orange/ethidium bromide. (D) control, (E) and (F) treated with CdNPs and CdNPs@BSA, respectively. (Down) DNA ladder on agarose gel electrophoresis. MDA-MB-231 Cells were treated with LD50 concentrations of CdNPs and CdNPs@BSA for 24 h and DNA fragmentation was analyzed. Lane M: 1 kB ladder, lane 1: control, lane 2: CdNPs, lane 3: CdNPs@BSA.
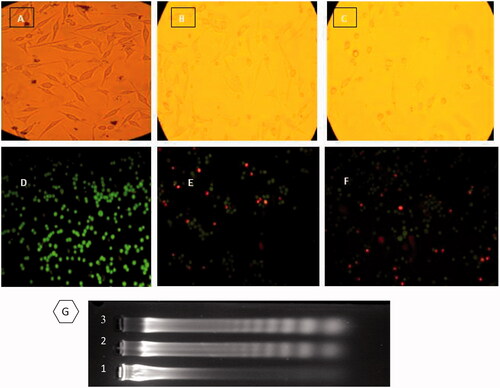
Figure 6. (A–C) Two-dimensional contour density plots of MDA-MB-231 cells obtained by flow cytometry based assays. A: Control, B-C: Treated cells with CdNPs and CdNPs@BSA, respectively. Cell necrosis and apoptosis were measured by using 7-AAD and Annexin-V dyes. (D–F) Flow cytometry based assay of cell cycle. D: Control, E–F: Treated cells with CdNPs and CdNPs@BSA, respectively.
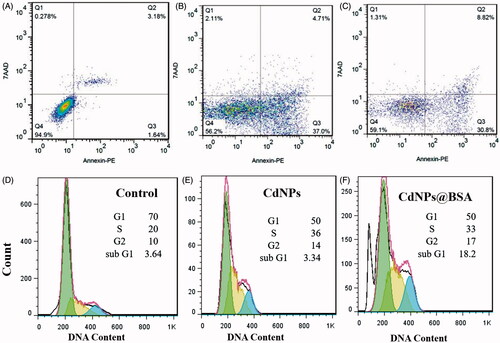
Table 1. Quantified flow cytometry analysis of PE annexin V and 7-AAD stained MDA-MB-231 cells after 24 h treatment with CdNPs and CdNPs@BSA.
Figure 7. Two-dimensional contour density plots of WBCs cells obtained by flow cytometry-based assays. (A) Control, (B) Treated cells with CdNPs@BSA. Cell necrosis and apoptosis were measured by using 7-AAD and Annexin-V dyes.
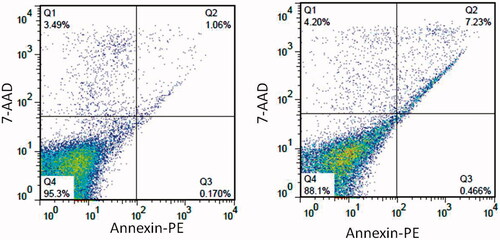
Table 2. Quantified flow cytometric analysis of PE annexin V and 7-AAD stained WBCs cells after 24 h treatment with CdNPs@BSA.