Figures & data
Figure 1. Characterization of PVA and PVA-H microspheres. (a) The shape (300×) and porous structure (10,000×) micrographs of PVA and PVA-H microspheres by SEM. (b) Bright field, fluorescent and merge photographs of PVA and PVA-H/FITC. For each panel, images from left to right show bright field, fluorescent photographs, and merge of two images, respectively. (c) The XPS wide spectrum of PVA-H and the fitted peak analysis of the XPS for N 1 s and S 2p of PVA-H.
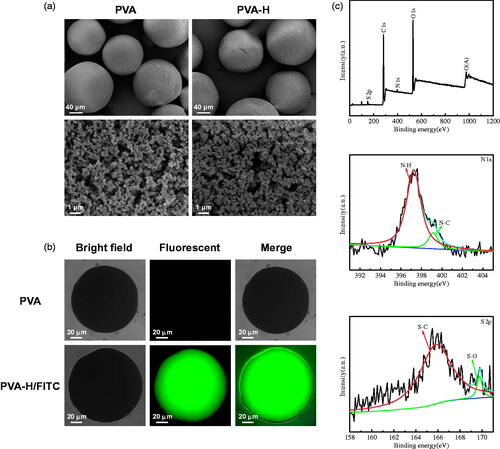
Figure 2. The adsorption characteristics of PVA-H for VEGF, TGF-β and TNF-α in human plasma. (a) Non-linear fitting for the adsorption kinetic data of VEGF onto the two adsorbents in human plasma (T = 37 °C, CVEGF = 1335.21 ± 3.92 pg/mL). (b) Effects of temperature on adsorption of VEGF (t = 2 h, CVEGF = 1152.32 ± 4.23 pg/mL). (c) Non-linear fitting for the adsorption kinetic data of TGF-β onto the two adsorbents in human plasma (T = 37 °C, CTGF-β = 1048 ± 1.24 pg/mL). (d) Effects of temperature on adsorption of TGF-β (t = 2 h, CTGF-β = 1121 ± 2.34 pg/mL). (e) Non-linear fitting for the adsorption kinetic data of TNF-α onto the two adsorbents in human plasma (T = 37 °C, CTNF-α = 1069 ± 3.24 pg/mL). (f) Effects of temperature on adsorption of TNF-α (t = 2 h, CTNF-α = 968 ± 4.25 pg/mL). The plasma to adsorbent ratio was 20 and all values are expressed as mean ± SD (n = 3). ns: not significant, **p < .01, ***p < .001, ****p < .0001.
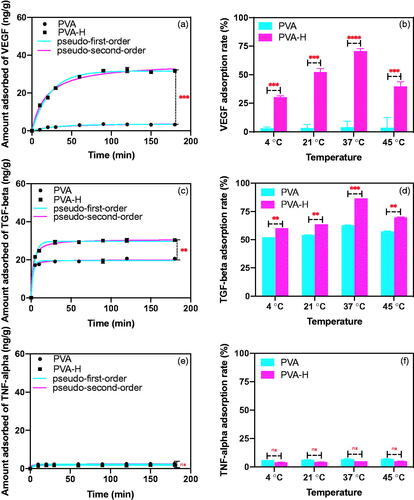
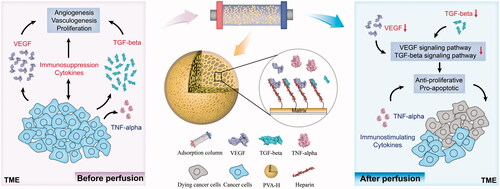
Figure 3. Targeting regulation of the TME in SHZ-88 cells by PVA-H microspheres. (a) Schematic illustration of PVA-H microspheres dynamic targeting experiment. (b) Adsorption rate of different cytokines in the culture supernatants after perfusion through PVA and PVA-H microspheres (T = 37 °C, CVEGF = 490.83 ± 5.32 pg/mL, CTGF-β = 345.92 ± 6.32 pg/mL, CTNF-α = 4.12 ± 1.24 pg/mL the plasma to adsorbent ratio was 20). (c) CCK-8 experiments showed that perfusion through PVA-H microspheres attenuated the proliferative capacity of SHZ-88 cells. All values are expressed as mean ± SD (n = 6). ns: not significant, **p < .01, ***p < .001, ****p < .0001.
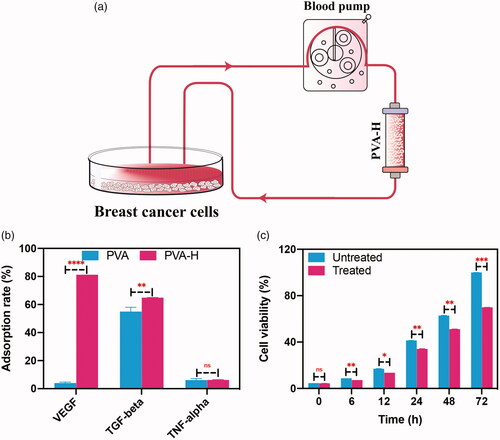
Figure 4. Annexin V–FITC and PI assay. Effect of perfusion through PVA-H microspheres on cell apoptosis. All values are expressed as mean ± SD (n = 3).
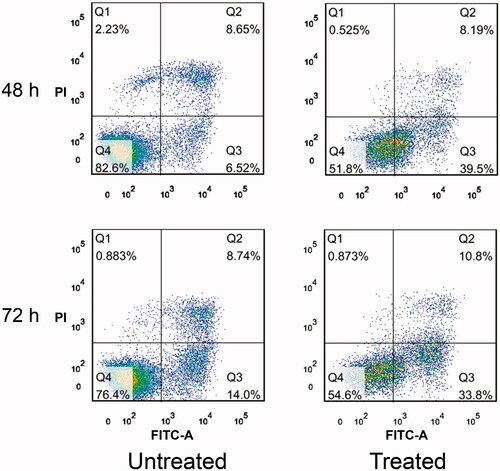
Figure 5. Mechanism of the anti-proliferative and pro-apoptotic effects on breast cancer cells. SHZ-88 cells were cultured for 12 h and then exposed to supernatants treated or not treated by perfusion through PVA-H microsphere columns. (n = 4). (a) Genes involved in positive and negative regulation from the genes involved in proliferation and apoptosis. (Left: Genes associated with positive and negative regulation of cell proliferation; Right: Genes involved in positive and negative regulation of apoptosis). (b) Functionally significantly related genes involved in the proliferative and apoptotic functions (Left: proliferation-related genes; Right: apoptosis-related genes).
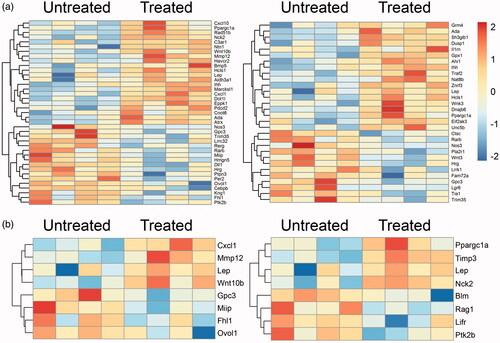
Figure 6. KEGG pathway enrichment analysis of the differentially expressed genes (Reds are up-regulated, petunias are down-regulated).
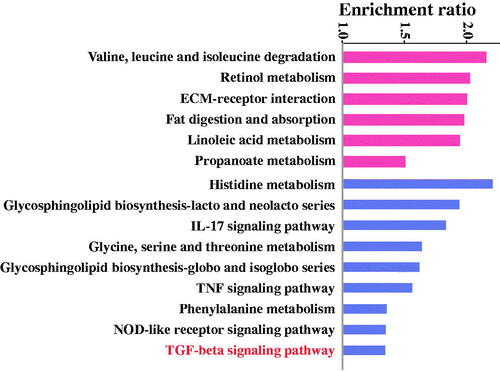
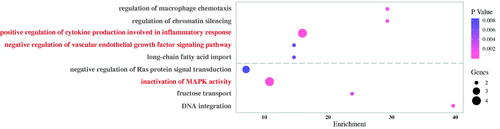
Figure 7. GO enrichment analysis of the differentially expressed genes (GO names above the imaginary line are up-regulated, GO names under the imaginary line are down-regulated).
Supplemental Material
Download MS Word (13.6 MB)Data availability statement
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary materials.