Figures & data
Figure 1. The molecular structure of SARS-CoV-2 S protein. (A) The schematic diagram of the domain structure of SARS-CoV-2 S protein. NTD, N-terminal domain (yellow). RBD, receptor-binding domain (cyan). S1/S2, S1/S2 furin cleavage site. S2’, S2’ cleavage site R815 (blue spheres). FP, fusion peptide (bright purple). HR1, heptad repeat 1 (green). HR2, heptad repeat 2 (brown). (B) The trimeric S protein ectodomain. (C) The prefusion (left) and postfusion (right) structures of the trimeric S2 subunits. HR2 was not seen in the crystal structure of the trimeric S protein ectodomain (PDB ID: 6XR8).
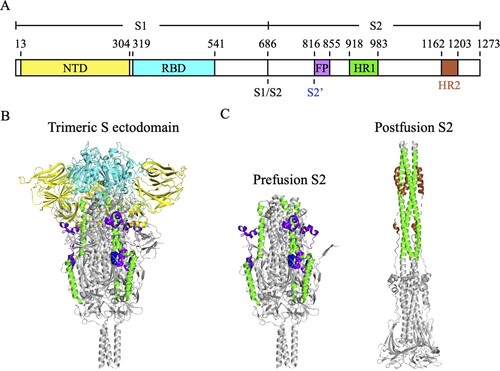
Figure 2. An immunodominant S2 peptide recognized by various S2 stem helix-specific antibodies. (A) Sequence alignment of the S2 stem helix regions (S1141-1160) of SARS-CoV-2 VOCs, including Wuhan-Hu-1, Alpha, Beta, Gamma, Epsilon, Delta, and Omicron strains. (B) Sequence alignment of the S2 stem helix regions of SARS-CoV-2, SARS-CoV, MERS-CoV, HCoV-OC43, HCoV-HKU1, HCoV-NL63, and HCoV-229E. S1141-1160 is shown as SARS-CoV-2 numbering. Residues identical to the sequence of SARS-CoV-2 S2 stem helix are marked with black backgrounds. (C) The key binding epitopes of the S2 stem helix-specific antibodies. Hydrophobic, negatively charged, and positively charged residues of S1141-1160 are coloured in grey, magenta, and brown, respectively. Residues recognized by S2P6 and 1249A8 that broadly neutralize SARS-CoV-2, SARS-CoV, and MERS-CoV are coloured in cyan and marine blue, respectively. Residues recognized by CC40.8, CC68.109, and CC99.103 that neutralize SARS-CoV-2 and SARS-CoV but not MERS-CoV are coloured yellow. Residues recognized by WS6 that neutralizes SARS-CoV-2 and SARS-CoV but not MERS-CoV are coloured in orange. Residues E1144, F1148, L1152, and F1156 recognized by S2-4D, S2-5D, S2-8D, and S2-4A that neutralize SARS-CoV-2 are coloured in pink. Residues recognized by B6, IgG22, 1.6C7, and 28D9 that only neutralize MERS-CoV but not SARS-CoV-2 are coloured in green, purple, and light purple, respectively.

Table 1. The SARS-CoV-2 S2 stem helix-specific neutralizing antibodies.
Figure 3. Superimposition of S2 stem helix-specific antibodies upon binding with their epitopes. (A) The Fabs of S2P6 (cyan, PDB: 7RNJ), CC40.8 (yellow, PDB: 7SJS), WS6 (orange, PDB: 7TCQ), B6 (green, PDB: 7M53), IgG22 (purple, PDB: 7S3N), and CV3-25 (magenta, PDB: 7RAQ) are superimposed with the prefusion S trimer (grey, PDB: 6XR8). (B) Alignment of the Fabs of S2P6, CC40.8, WS6, B6, IgG22, and CV3-25 upon binding with residues 1141-1162 of the S2 stem helix. Superimposition of the Fabs of S2P6 and B6 (C), CC40.8 and WS6 (D), or B6 and IgG22 (E) for comparison.
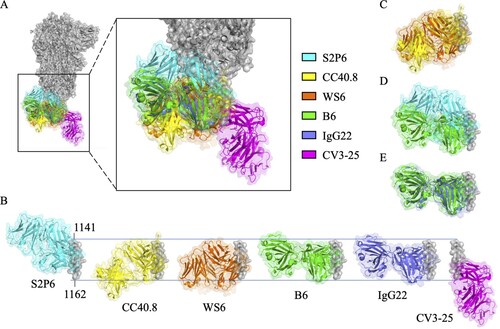
Table 2. The SARS-CoV-2 S2’/FP-specific neutralizing antibodies.
Figure 4. Sequence alignment of the partial FP residues recognized by S2’/FP-specific antibodies. (A) Sequence alignment of the partial S2’/FP regions (S815-830) of SARS-CoV-2 VOCs, including Wuhan-Hu-1, Alpha, Beta, Gamma, Epsilon, Delta, and Omicron strains. (B) Sequence alignment of the partial S2’/FP regions of SARS-CoV-2, SARS-CoV, MERS-CoV, HCoV-OC43, HCoV-HKU1, HCoV-NL63, and HCoV-229E. S815-830 is shown as SARS-CoV-2 numbering. Residues identical to the S2’/FP sequence of SARS-CoV-2 are marked with black backgrounds. (C) The key residues recognized by 76E1, COV44-62, COV44-79, and VN01H1 that can neutralize both of α- and β-CoVs are coloured in cyan, yellow, magenta, and green, respectively. The key residues recognized by C77G12 that can only neutralize β-CoVs are coloured in orange.

Figure 5. Superimposition of S2’/FP-specific antibodies upon binding with their epitopes. The Fabs of 76E1 (cyan, PDB: 7X9E), COV44-62 (yellow, PDB: 8D36), COV44-79 (magenta, PDB: 8DAO), VN01H1 (green, PDB: 7SKZ), and C77G12 (orange, PDB: 7U0A) are superimposed with the FP (bright purple) of SARS-CoV-2 S protein. N’ and C’ stand for the N- and C-termini of FP.
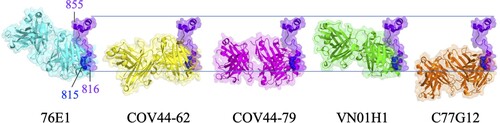
Figure 6. Sequence alignment of the partial HR2 residues recognized by hMab5.17. (A) Sequence alignment of the partial HR2 residues of SARS-CoV-2, SARS-CoV, MERS-CoV, HCoV-OC43, HCoV-HKU1, HCoV-NL63, and HCoV-229E. S1163-1176 is shown as SARS-CoV-2 numbering. Residues identical to the sequence of SARS-CoV-2 HR2 are marked with black backgrounds. (B) Sequence alignment of the partial HR2 residues (S1163-1176) of SARS-CoV-2 VOCs, including Wuhan-Hu-1, Alpha, Beta, Gamma, Epsilon, Delta, and Omicron strains. F1176 in Gamma strain is marked with black background. (C) The binding epitope of hMab5.17 on the SARS-CoV-2 S protein is coloured in grey.

