Figures & data
Table 1. Sample collection distribution and classification.
Figure 1. Distribution of captured bat species in the sampled areas showing the results of coronavirus detection.
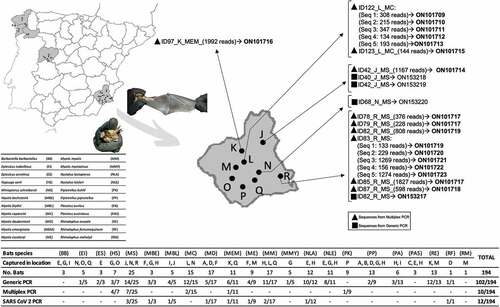
Table 2. Primers used for partial gene amplifications.
Figure 2. Phylogenetic tree generated for the 118 bp fragment from the coronavirus generic PCR (ORF1b region).
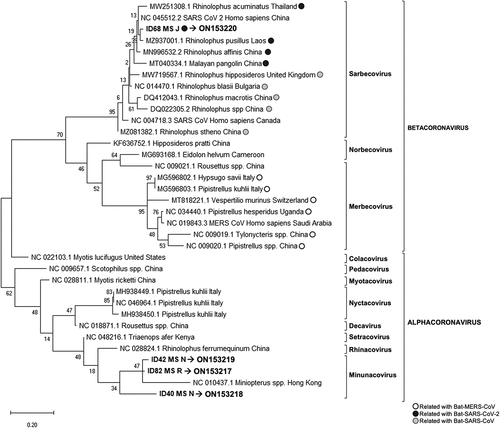
Figure 3. Phylogenetic tree generated for the 381 bp fragment from the coronavirus Multiplex-PCR (ORF1ab region).
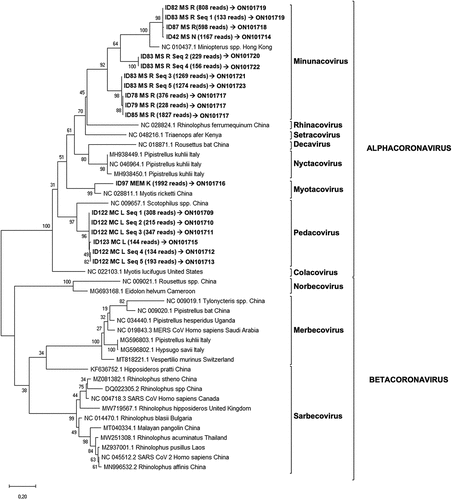
Table 3. NGS sequencing results.
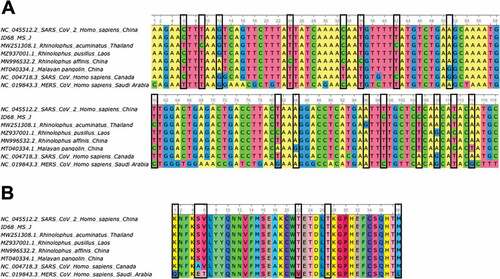
Figure 4. A) Alignment of partial ORF1b gene sequences (118 bp) of some bat-related SARS-CoV-2 (MN996532.2, MZ937001.1, MW251308.1) and pangolin (MT040334.1) coronavirus isolates, including the SARS-CoV-2 Wuhan-Hu-1 (NC_045512.2), SARS-CoV (NC_004718.3) and MERS-CoV (NC_019843.3) reference sequences and the sequence obtained in this study ID 68 (ON153220). Nucleotide changes between bat and pangolin isolates related to SARS-CoV-2 with the reference sequence for this virus are highlighted in black boxes. B) Amino acid alignment (39 aa) of partial ORF1b gene sequences (118bp) described in section A of this figure. Amino acid changes between bat and pangolin isolates related to SARS-CoV-2 with the reference sequence for SARS-CoV and MERS-CoV are highlighted in black boxes.