Figures & data
Figure 2. Displacement ellipsoids (50% probability) plot of the asymmetric unit depicting the molecular structures of the two independent [Ga2Br2]2− anions and the three independent [Ph3PH]+ cations (Ga1 with site symmetry ; Ga2,3 with site symmetry 3). False colours indicate symmetry-related atoms (sym codes on diagram). A symmetry-equivalent P1 cation additional to the asymmetric unit is shown in light grey (‘‘). Phenyl ring T-interactions from C–H to C are shown with dashed blue lines once for each symmetry-replicated interaction. H atoms on carbon are omitted, as are atom labels for symmetry related atoms.
![Figure 2. Displacement ellipsoids (50% probability) plot of the asymmetric unit depicting the molecular structures of the two independent [Ga2Br2]2− anions and the three independent [Ph3PH]+ cations (Ga1 with site symmetry 3¯; Ga2,3 with site symmetry 3). False colours indicate symmetry-related atoms (sym codes on diagram). A symmetry-equivalent P1 cation additional to the asymmetric unit is shown in light grey (‘‘). Phenyl ring T-interactions from C–H to C are shown with dashed blue lines once for each symmetry-replicated interaction. H atoms on carbon are omitted, as are atom labels for symmetry related atoms.](/cms/asset/417168bd-cd0e-4dc2-8c9f-f6061cb11d4c/oach_a_1273065_f0002_oc.gif)
Table 1. Selected interatomic distances, angles and torsions in the crystal structure of the title compound
Figure 3. Packing diagram of the structure of the title compound in P. H atoms on the phenyl rings are omitted for clarity, as are all atoms along the front (1 1 0) unit cell edge. The phosphonium ions are rendered as rods, whereas the hexabromodigallate ions are rendered as “space filling” spheres.
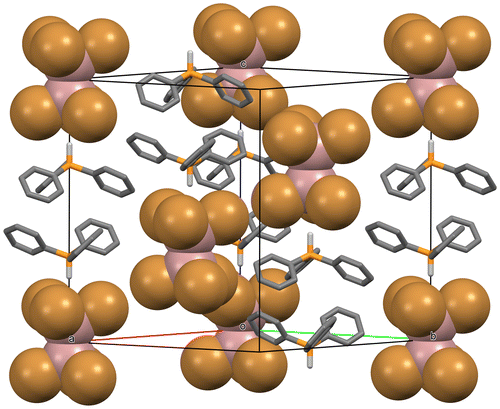
Figure 4. Overlaid unit cells of the title structure in P with the alternate refinement in R
with apparent disorder as in (Khan et al., Citation1986), drawn as 30% displacement ellipsoids. The
body diagonal of the rhombohedral cell is aligned with the 3 axis of the trigonal cell along (⅔ ⅓ 0).
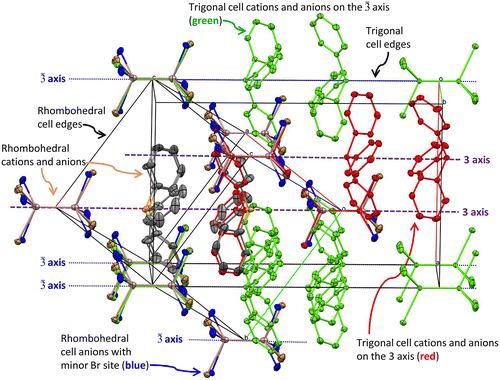
Figure 5. Representation of part of the crystal structure of Li2[Ga2Br6]; there is a mirror plane through Br11, Ga1, Ga2, Br22. Li+ ions coordinated to Br22 and all remaining bromine atoms completing the coordination spheres of Li1,2 have been omitted for clarity. The data is taken from Hönle and Simon (Citation1986).
![Figure 5. Representation of part of the crystal structure of Li2[Ga2Br6]; there is a mirror plane through Br11, Ga1, Ga2, Br22. Li+ ions coordinated to Br22 and all remaining bromine atoms completing the coordination spheres of Li1,2 have been omitted for clarity. The data is taken from Hönle and Simon (Citation1986).](/cms/asset/4aaf6b58-2a65-4eec-8cb2-bdf2d1d06092/oach_a_1273065_f0005_oc.gif)
Figure 6. Conformational energy profile for the title anion computed at the B3LYP/6–311+G(fd,) level of theory for gas-phase [Ga2Cl6]2– and [Ga2Br6]2– ions. The calculations were undertaken in 10° steps in the X–Ga–Ga–X torsion angle starting from 63.3°.
![Figure 6. Conformational energy profile for the title anion computed at the B3LYP/6–311+G(fd,) level of theory for gas-phase [Ga2Cl6]2– and [Ga2Br6]2– ions. The calculations were undertaken in 10° steps in the X–Ga–Ga–X torsion angle starting from 63.3°.](/cms/asset/106baf87-06c8-48e9-b3c2-dcc357698d7a/oach_a_1273065_f0006_oc.gif)

![Figure 1. Disordered [Ga2Br6]2− anion in the refinement model for FUPSIS.](/cms/asset/d8698906-9190-4b9f-ac1d-290bbbbc3927/oach_a_1273065_f0001_b.gif)